Arthur Viode retweetledi
Arthur Viode
13 posts

Arthur Viode retweetledi

Predictive gene expression signature diagnoses neonatal sepsis before clinical presentation - eBioMedicine thelancet.com/journals/ebiom…
English
Arthur Viode retweetledi

New large-scale plasma proteome profiling study, 1100+ samples, 6Tb of data, 2500+ proteins quantified. Analysis using Bruker DDA-PASEF. I actually think DDA MS1 LFQ is a solid strategy and even has some advantages, even in what seems to be the era of DIA. science.org/doi/10.1126/sc…
English
Arthur Viode retweetledi

Want reliable quantities? Brilliant students in my lab @ZiskaKistner @JustusGrossmann figured out how. We present QuantUMS, an ML-based algorithm for step-change better quantitation in proteomics and statistical confidence in individual quantities biorxiv.org/content/10.110…
QuantUMS is integrated in DIA-NN (beta version referenced in preprint), but we also plan to release it as an open-source tool, for use on various kinds of data, not just DIA.
We further anticipate significant gains to be achieved by integrating the accuracy metric reported by QuantUMS with existing packages for statistical analysis of proteomics data.
Some benchmarks are in the twitter thread below.

English
Arthur Viode retweetledi

Check out our perspective on MS-based body fluid #proteomics in @molcellprot.
We discuss how recent and cutting-edge advances overcome long-standing challenges and bring the field to an exciting turning point.
Great thanks to Jakob Bader and @labs_mann
mcponline.org/article/S1535-…
English
Arthur Viode retweetledi

Meta-analysis of published cerebrospinal fluid proteomics data identifies and validates metabolic enzyme panel as Alzheimer’s disease biomarkers cell.com/cell-reports-m…
Română
Arthur Viode retweetledi

'Using mass spectrometry to validate mouse models of #tauopathy'
Yan Yan & Casey N. Cook @MayoClinicNeuro #AlzheimersDisease #FrontotemporalDementia
bit.ly/3ZHqqbx

English
Arthur Viode retweetledi

Our new study in @NatureComms evaluates tools to detect longitudinal differential expression in #proteomics data and introduces a robust approach RolDE for longitudinal #biomarkerdiscovery. @BioscienceTurku @UniTurku @SuomenAkatemia @InFLAMES_Health rdcu.be/c15A8
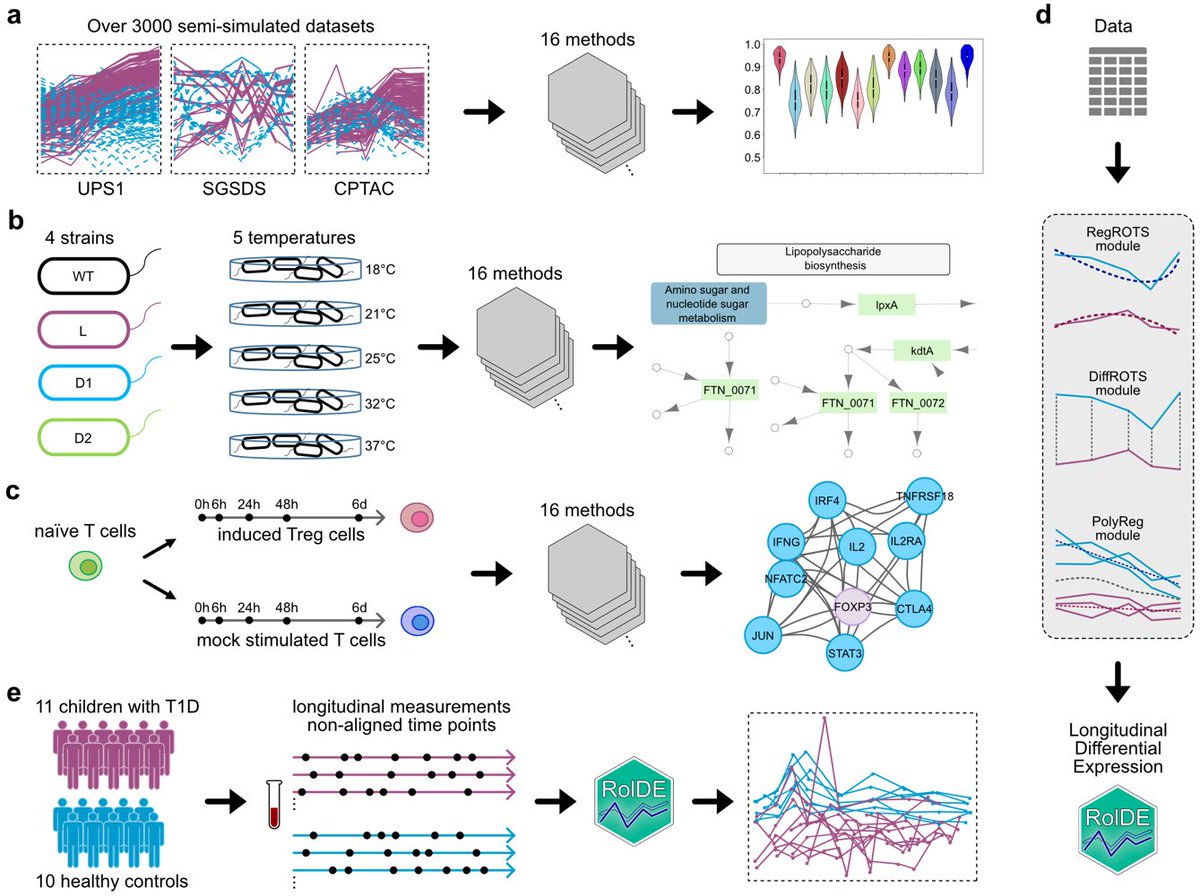
English

Just introduced a game-changing plasma depletion method for deep and high throughput protein profiling!
@bruker @EvosepBio @ScienceAdvances #proteomics #massspectrometry #innovation #clinicalresearch #COVID19 #HarvardMed
science.org/doi/10.1126/sc…
English
Arthur Viode retweetledi

The Hanno Steen lab at Boston Children's' Hospital optimized #FragPipe for a high-performance computing environment, and applied it to analyze 3348 timsTOF plasma samples (6.4 Tb) from a longitudinal COVID-19 patients study. pubs.acs.org/doi/full/10.10…

English
Arthur Viode retweetledi

The @steen_lab presents a Fragpipe-based parallelization strategy for analysis of LC-MS proteomics data. Their strategy reduced run time by 90% & enables them to map the proteomes of 1000s of samples! pubs.acs.org/doi/10.1021/ac…

English
Arthur Viode retweetledi

'Common mouse models of #tauopathy reflect early but not late human disease'
@steen_lab @ionicwoman @harvardmed #AlzheimersDisease #proteomics
bit.ly/3Y21GuI

English
Arthur Viode retweetledi

Kathrin Wenger and our team asked, Why do drugs for Alzheimer’s Disease work in animal models, but often fail in clinical trials?
…arneurodegeneration.biomedcentral.com/articles/10.11…
#AlzheimersDisease #tauopathy #proteomics #HarvardMed #neurodegeneration #dementia

English
