
Dan Polasky
48 posts

@DanPolasky
Research Investigator @ nesvilab - University of Michigan | mass spectrometry, data analysis and software, ion mobility, proteomics














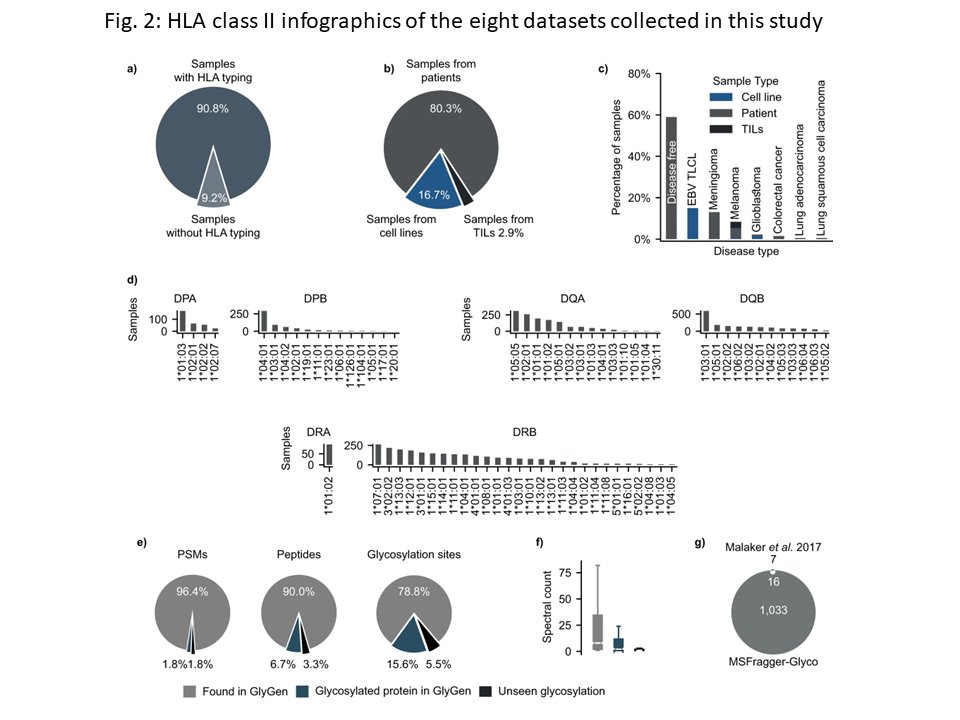
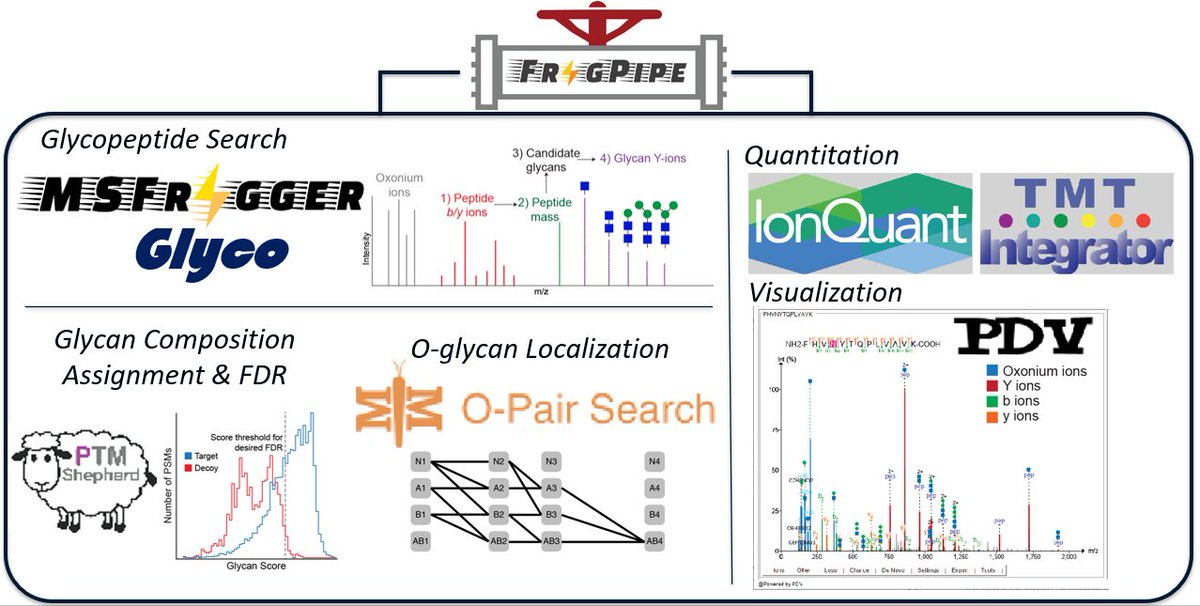
Unraveling the glycosylated immunopeptidome! Here we describe our HLA-Glyco workflow in #FragPipe, a web resource based on eight large mass spec based immunopeptidome datasets (over 3,400 HLA class II N-glycopeptides), and several interesting observations. nature.com/articles/s4146…







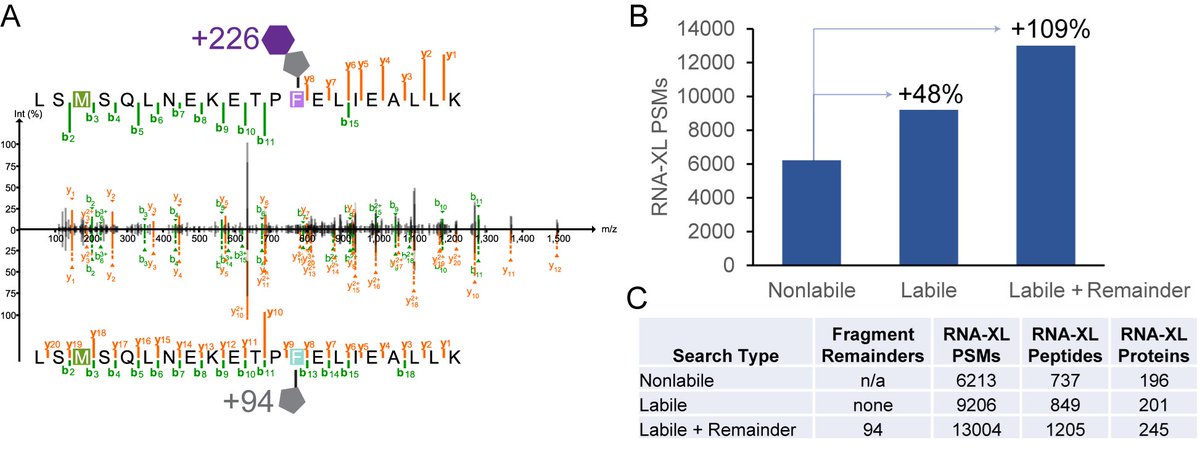







Over half of the peptides in the #immunopeptidome are still unidentified! Check out our new paper in @NatureBiotech where we develop PROMISE to identify modified HLA peptides and illuminate🔦 MS dark matter, revealing new classes of cancer antigens. nature.com/articles/s4158…🧵1/7

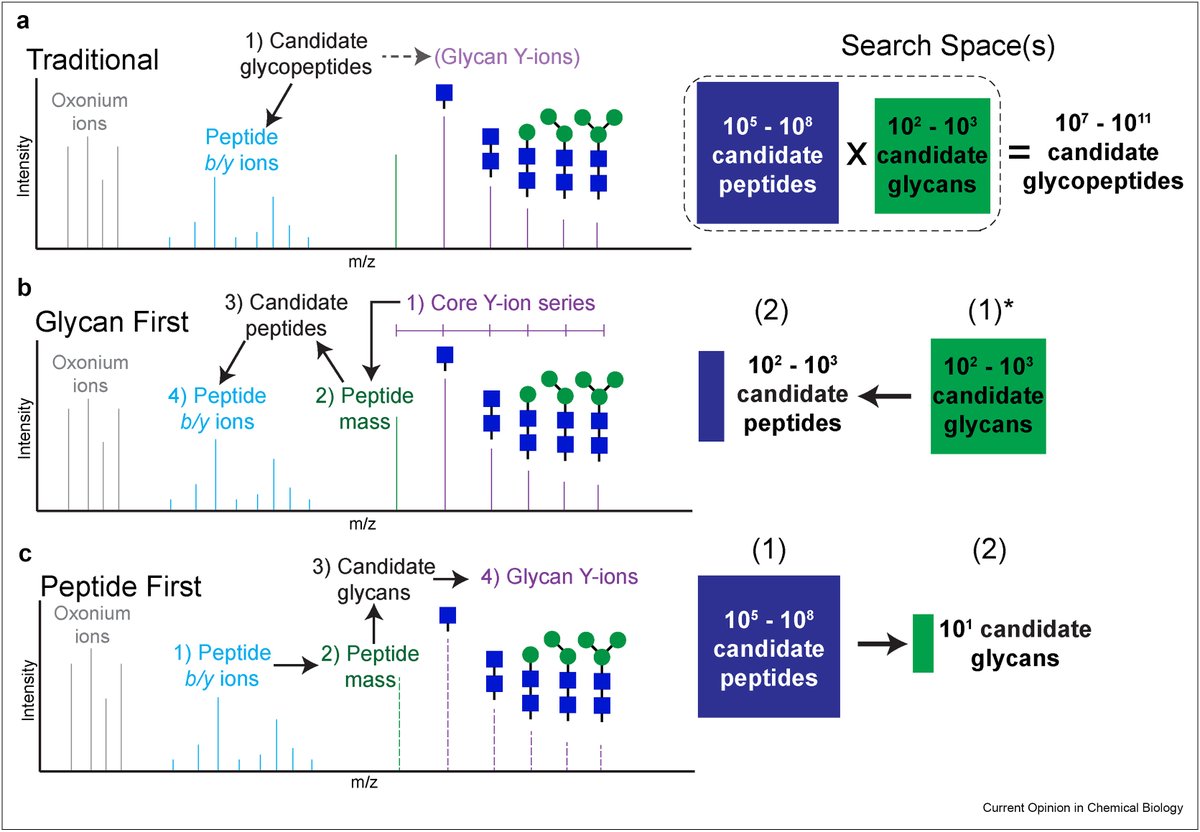
Our latest Primer summarizes glycoproteomic techniques and best practices for their use @kattesch @StacyMalaker @BagdonaiteIeva go.nature.com/3xQ84td
