
Ida Lindeman
55 posts

Ida Lindeman
@IdaLindeman
Postdoc at Oslo University Hospital Interested in autoimmunity, celiac disease, plasma cells, BCRs, IG polymorphisms and single-cell technologies.




Generation of circulating autoreactive pre-plasma cells fueled by naive B cells in celiac disease biorxiv.org/cgi/content/sh… #biorxiv_immuno


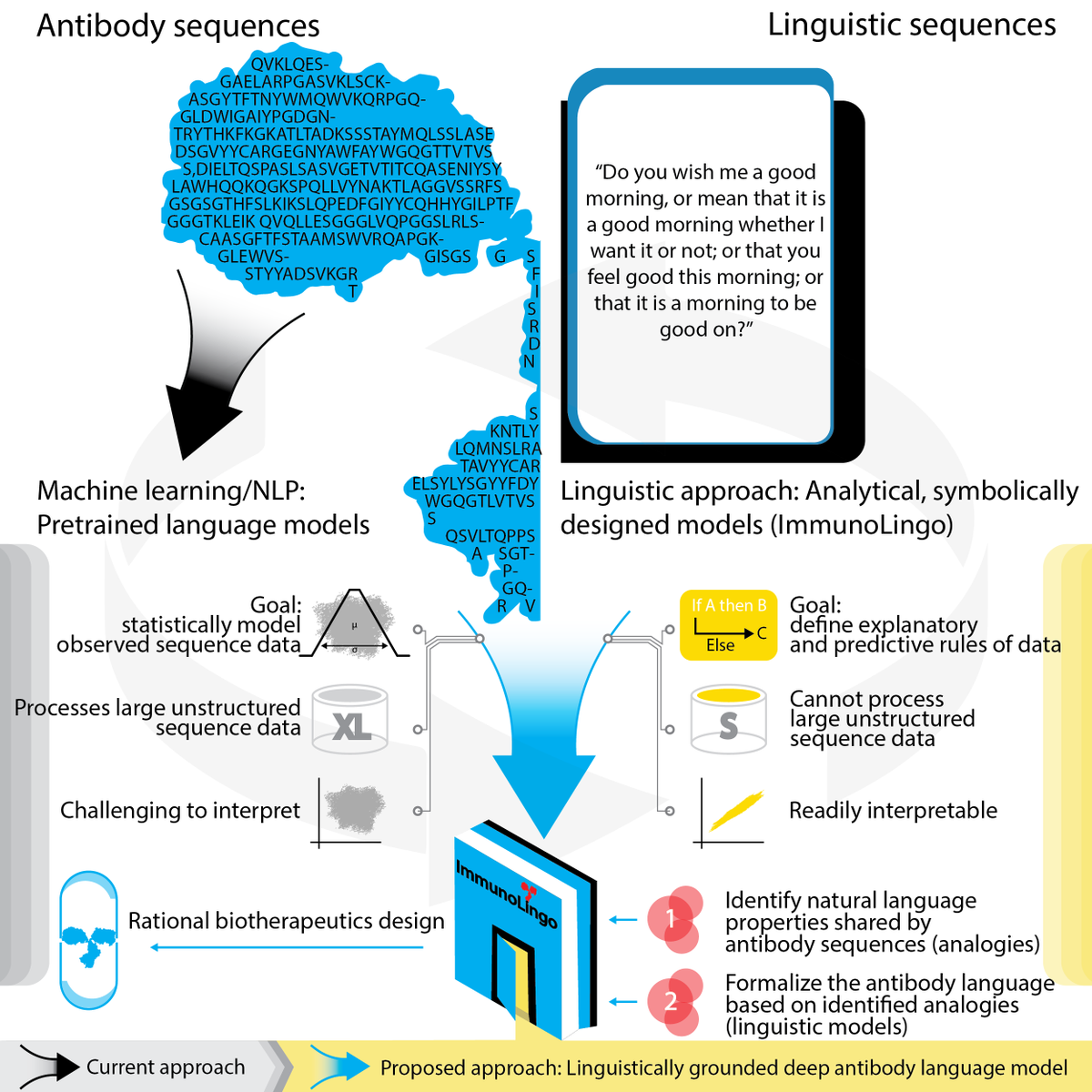
New preprint from the ImmunoLingo convergence environment: “ImmunoLingo: Linguistics-based formalization of the antibody language”. We asked: how to define the language of antibody sequences for more efficient antibody therapeutics design? 1/7 arxiv.org/abs/2209.12635





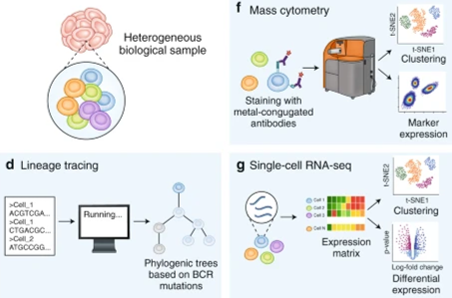

We are happy to report a dramatic speedup for one of the core computations for adaptive immune receptor repertoire (AIRR) analysis - the discovery and counting of receptors that overlap between repertoires! Check out our CompAIRR method: biorxiv.org/content/10.110… 1/7

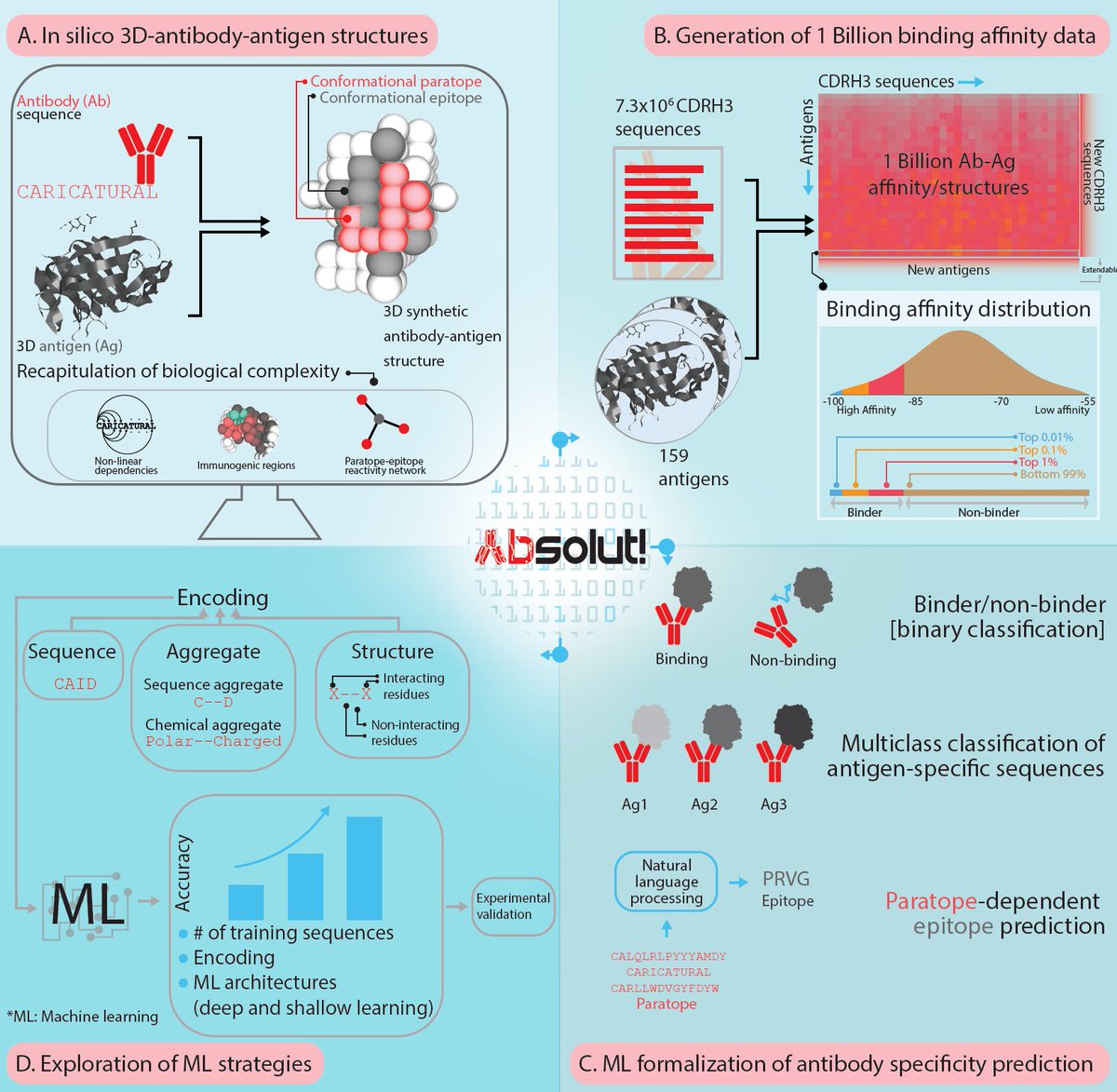



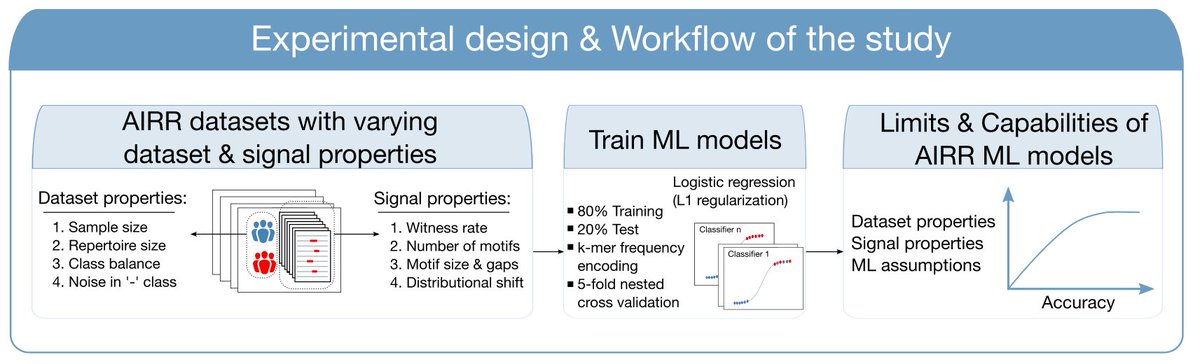






Very happy to finally release immuneML – an open-source python platform that streamlines development, application, and benchmarking of immune receptor machine learning for predicting immune status or antigen specificity (immuneml.uio.no). biorxiv.org/content/10.110… 1/10