Allon Klein Lab
69 posts


Allon Klein Lab
@KleinLabHMS
Klein lab @HMS_SysBio. Fate decisions, dev bio, regeneration, single cell genomics, math, data sharing, rubber ducks. Tweets from lab members. Like/RT ≠ endorse



I remember seeing the very first tests in this magical work… Amazing science from @KleinLabHMS Multistep genomics on single cells and live cultures in subnanoliter capsules | Science science.org/doi/10.1126/sc…

I am delighted to share that my main PhD project is finally out in Science Magazine! I'm honored to have worked with so many talented colleagues and mentors that guided this journey and provided me an opportunity to contribute research! science.org/doi/10.1126/sc…


How does a stem cell "decide" its fate? Development requires both reliability (consistent cell types) AND flexibility (diverse outcomes from identical progenitors). Cells achieve this by dynamically tuning deterministic drift and stochastic diffusion. New in @NatMachIntell: scDiffEq models state-dependent drift AND diffusion, improving fate prediction by ~8% over SOTA. scDiffEq also enables genome-wide in silico perturbation screens and reveals temporal gene dynamics. 🧵nature.com/articles/s4225…



❔What is special about an immune system that successfully attacks tumors, compared to one that doesn’t? Our new study in @CellCellPress led by @jgungabeesoon, @Nich and @Mate_Kiss_ aimed to address this question. A joint work with the great @KleinLabHMS authors.elsevier.com/a/1gqieL7PXmbtw


Want to apply single-cell multi-omic lineage tracing to study your favorite system in mice at defined time point and with superior lineage resolution? Jointly with @Li_Li_666 @SarahBowling15 @FD_Camargo, we present DARLIN, a superior lineage-tracing mouse biorxiv.org/content/10.110…

Come join my group at Westlake university in the beautiful city of Hangzhou, China to study development and cell differentiation with advanced (multi-omic) lineage tracing approaches and machine learning! shouwenwang-lab.com

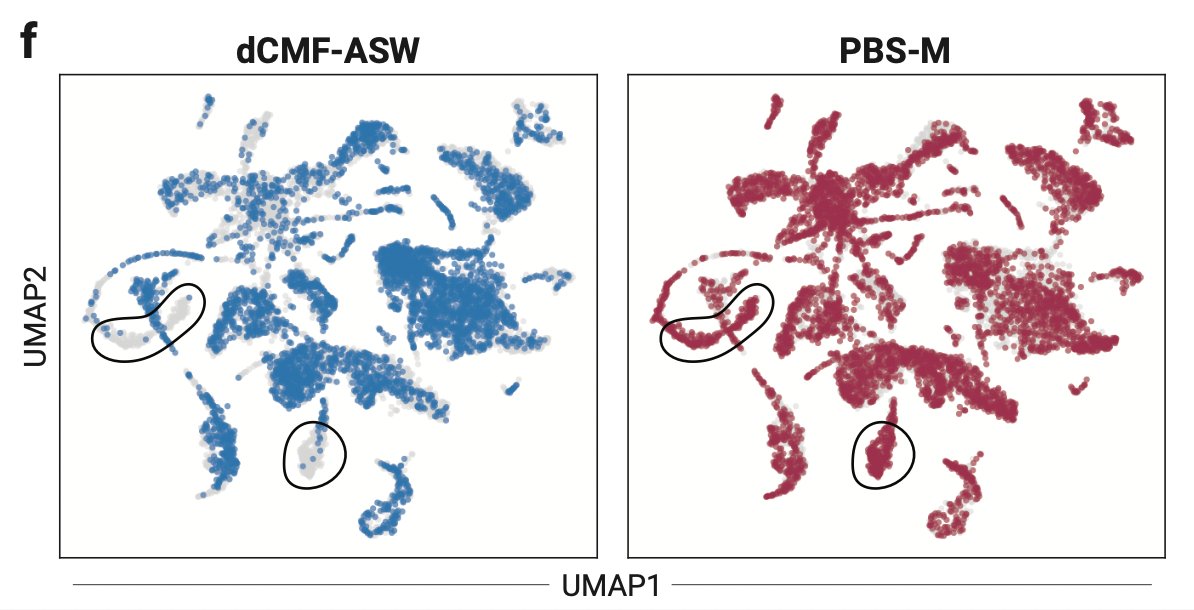
How do different signaling pathways alter tissue composition & state? Can we use signaling response phenotypes to predict aberrant signaling in disease? 🤔 We interrogate these in our new paper - biorxiv.org/content/10.110… A thread 1/9



I have accepted an offer for an assistant professor position at @Westlake_Uni starting in April 2023. Also, here is a recorded talk for my CoSpar paper in @NatureBiotech on integrating transcriptome and lineage information to learn early cell fate bias youtu.be/HrDQpW3kJFo.