Nick Monaco
25 posts

Nick Monaco
@NickMonaco99
Wellcome Optical Biology PhD @ UCL





You may have noticed some "holy $%@#" tweets on fly brain emulation. So is this a game-changer or a nothing-burger? Read on to find out...






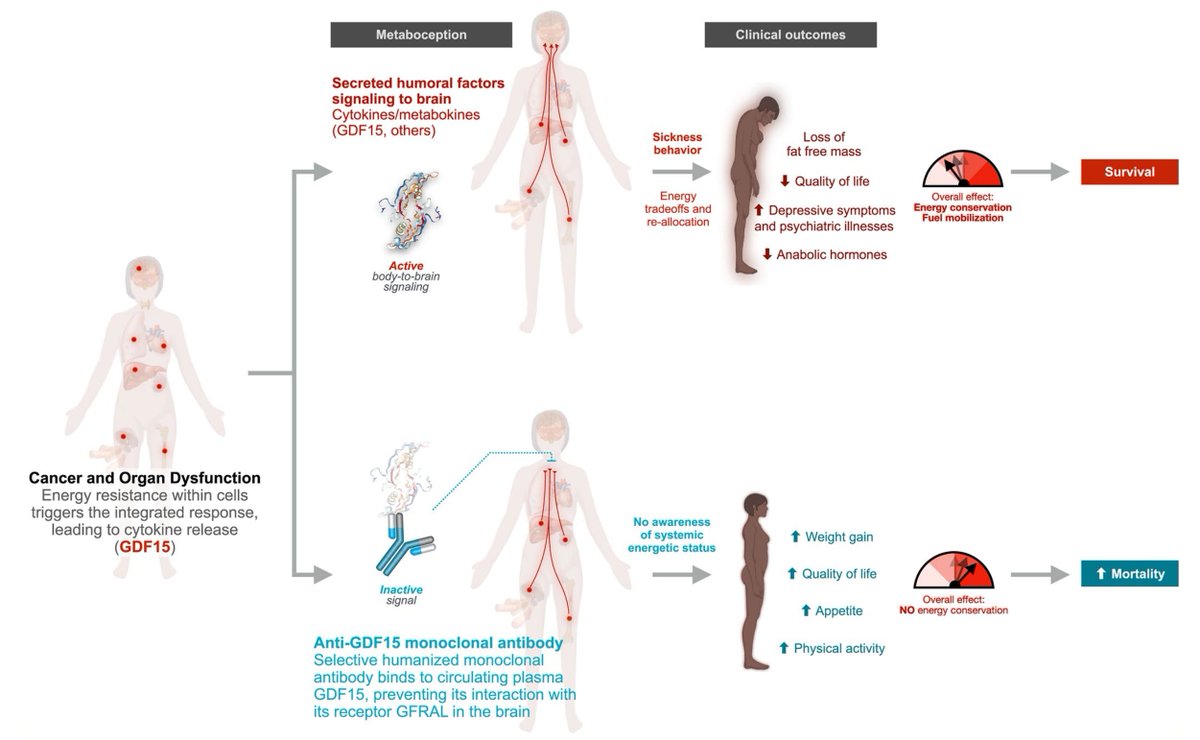
I’m going Founder Mode on my cancer. Below is Elliot Hershberg’s article about my cancer journey. It gave language to something I’d been doing instinctively over the past year: managing my health in Founder Mode. Manager mode assumes that existing systems will surface the best options. When I was first diagnosed with cancer in 2022, I delegated the crucial analyses and decisions about my care to others. In late 2024, when my cancer reappeared and my doctors told me I had exhausted the standard of care and there were no trials for my situation, I realized that assumption might, quite literally, kill me. Founder Mode was my only option. Founder Mode meant going deep on every diagnostic and treatment option. It meant assembling a team of physicians and scientists to work from first principles to understand what was possible beyond standard protocols. Together, we paved new roads to access the very cutting edge of science and technology. Today, thanks to the efforts of many people around the world and the support of my wife Karen, I currently have no evidence of disease. But my fight with cancer is far from over. My team and I continue to develop treatments and strategies in case it returns. More importantly, I now understand firsthand the challenges patients face in order to secure their own data and necessary treatments, particularly personalized medicines. I increasingly see my role as removing structural barriers—breaking down walls that prevent data, treatments, and technologies from flowing where they’re needed. One of the core principles of the first company I founded, GitLab, was radical transparency, and it’s a principle I am bringing to my cancer care. To that end, I am going to be sharing more about my experiences, my treatments, my data, and what I am building to make the path that I’ve been on easier for others to follow. Please subscribe to my mailing list on sytse.com to stay updated. Lastly, I want to thank those who have been on this journey with me. There have been too many to all thank here but I appreciate every one of you. I did want to mention Jacob Stern, Alfredo Gonzalez, and Jeremiah Wala; the amazing teams at Private Health Management (shoutout to Jenn and Eva) and Willy Hoos and Pathfinder Oncology; Nima Afshar and Private Medical; Sant Chawla and the Sarcoma Oncology Center; John Connolly and his team at the Parker Institute; Will Hudson at Baylor College of Medicine; Kamil Slowikowski for his work on osteosarc.com; and Jeff Tsao, Will Gibson, Ali Samiei, Scott McConnell and the rest of the team at the Briger Foundation for Oncology Research.



Two flagship papers from the International Brain Laboratory, now out in @Nature: 🧠 Brain-wide map of neural activity during complex behaviour → doi.org/10.1038/s41586… 🧠 Brain-wide representations of prior information in mouse decision-making → doi.org/10.1038/s41586… +





Today was my last lecture for @MIT #ComputationalBiology: #Genomes, #Networks, #Evolution, #Health. I recorded each and immediately posted online here: youtube.com/playlist?list=… Please do share, and let me know which topics need more explanations, clarifications, and corrections!

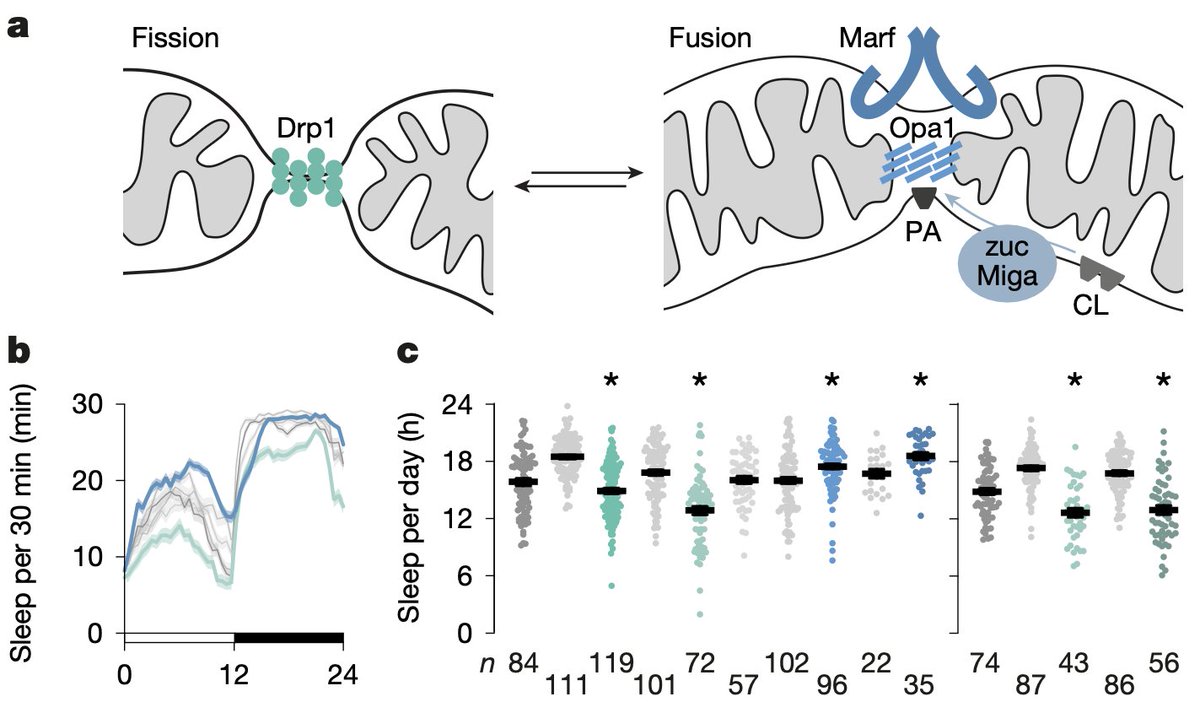



1/ 🚨 Our new paper is online in @Nature ! As its first author, I’m enthusiastic to finally share it with you all!🎉 🧠 We discovered a mechanistic link between cellular energy metabolism and the control of the need to sleep 💤 👉 nature.com/articles/s4158… @OxfordDPAG 🧵👇