
Zutao Yu
136 posts

Zutao Yu
@ZutaoYu
I am doing research in the Balasubramanian lab at Cambridge University. Working on nucleic acid Chemical biology.
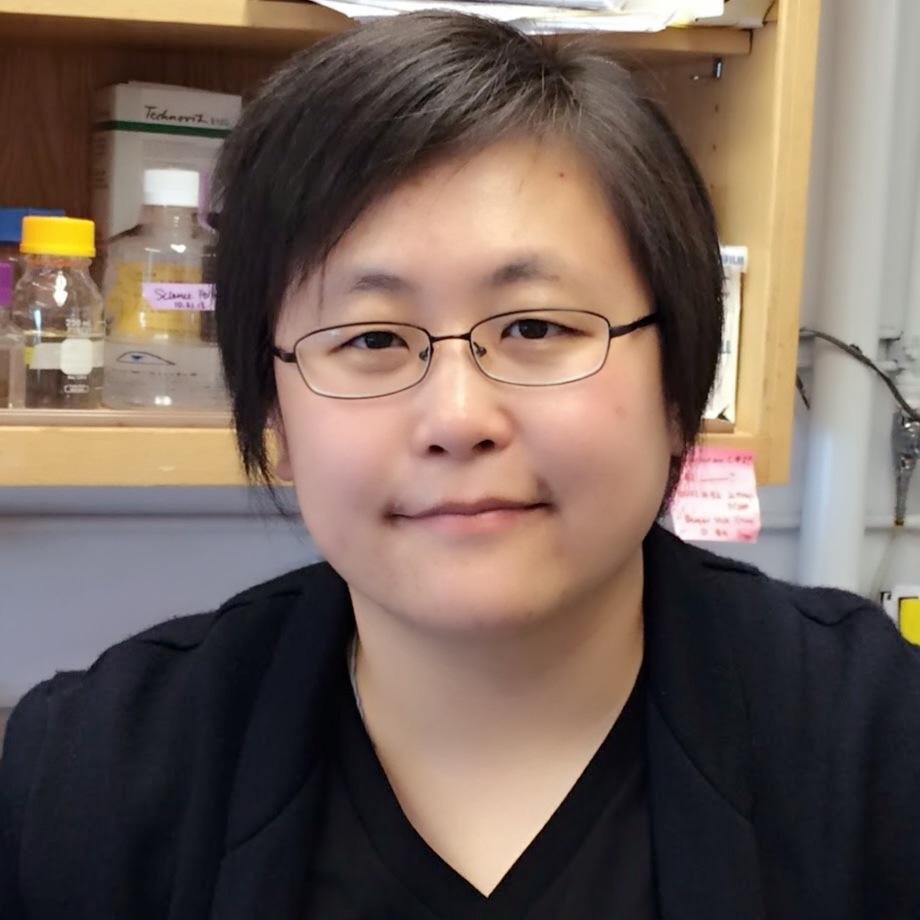



A true “lab” story in full bloom: Jonathan D. G. Jones, Wolf Prize Laureate in Agriculture 2025, and Caroline Dean, Wolf Prize Laureate in Agriculture 2020, are more than just partners in life — they’re also partners in discovery. Jonathan’s pioneering work revealed how plants defend themselves from disease, offering new ways to protect global crops. Caroline uncovered how plants use cold to control flowering time, a breakthrough in understanding how nature adapts to climate and how they “remember” winter. Two brilliant minds, one shared passion for plant science — and now, two Wolf Prizes under one roof. #wolfprize #wolfprizelaureate #agriculture @CarolineDeanLab @jonathandgjones

Please RETWEET! 2 postdocs (one chemical biologist, one RNA biologist) and multiple PhD positions are available at #KwokLab, City University of Hong Kong. We work on #RNA, #Gquadruplex, #aptamer , #peptide, #Biosensor and more! See kitkwok.com . PM if interested.








Engineering antisense oligonucleotides for targeted mRNA degradation through lysosomal trafficking ift.tt/jU35D49 #chemrxiv_biochem







We have dramatically increased the activity of splice switching *and* RNase H-active antisense oligonucleotide #ASO therapeutics by conjugating a nuclear importer! Preprint led by @disha_kashyap_ @milnetom68 @chem_in_cells @OxfordChemistry @UCLChemistry chemrxiv.org/engage/chemrxi…



Synthesizing RNA can be as fun as LEGO puzzles! This work is co-led by my incredible colleagues @LDangliang and @AbhiAditham . Huge thanks to my supervisor @WangXiaoLab for such amazing ideas. We are also grateful for the editor, reviewers, and all collaborators along the way.





Thrilled to share new @NatureNano paper by @FabriniGiacomo introducing genetically encoded condensate-forming RNA nanostructures🧬 doi.org/10.1038/s41565… ..and the great companion @NatureComms paper by @DrJMStewart @FrancoLabUCLA @pwkrpwkr doi.org/10.1038/s41467…

Transcriptome-Wide, Unbiased Profiling of Ribonuclease Targeting Chimeras | Journal of the American Chemical Society @disney_lab @UFScripps @scrippsresearch #Transcriptome #Profiling #Ribonuclease #Chimeras pubs.acs.org/doi/10.1021/ja…




In our new review paper, @ywan_wan and I focused on the functional importance of RNA structure in gene regulation. Many thanks to Yue and Xinang! I really enjoy the writing process. nature.com/articles/s4158…


