Nathanael Rollins
76 posts


@_nathanrollins
learning from evolution + synthesizing biology https://t.co/4R6uWsHqaL



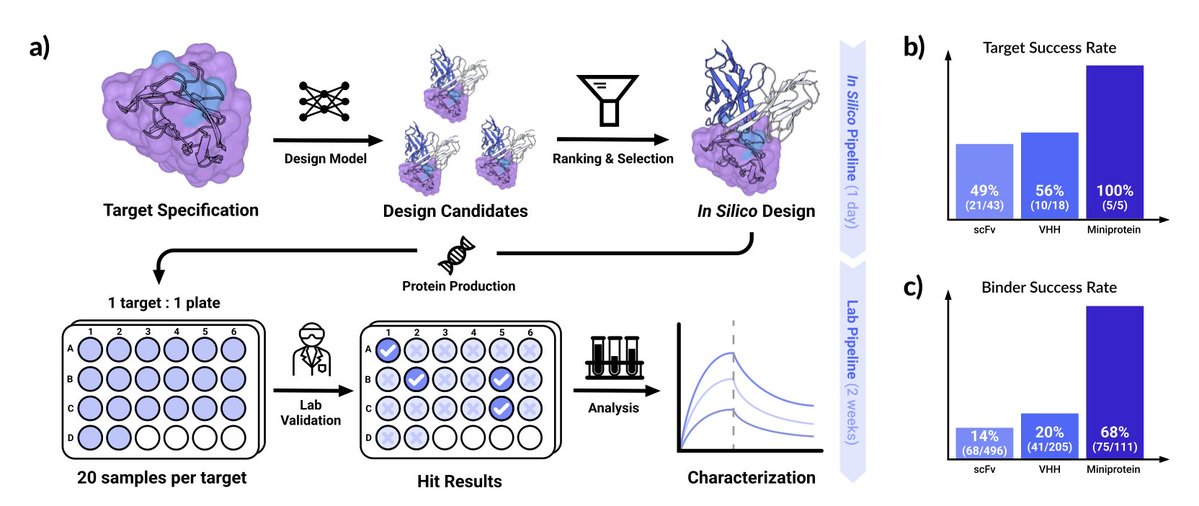
We’re excited to introduce Chai-2, a major breakthrough in molecular design. Chai-2 enables zero-shot antibody discovery in a 24-well plate, exceeding previous SOTA by >100x. Thread👇
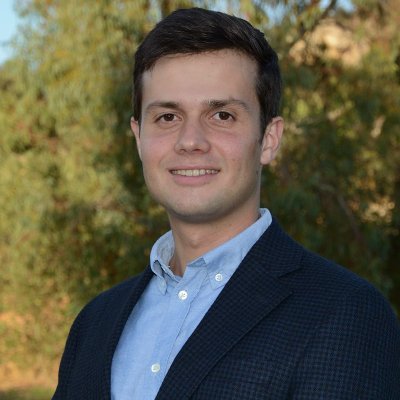





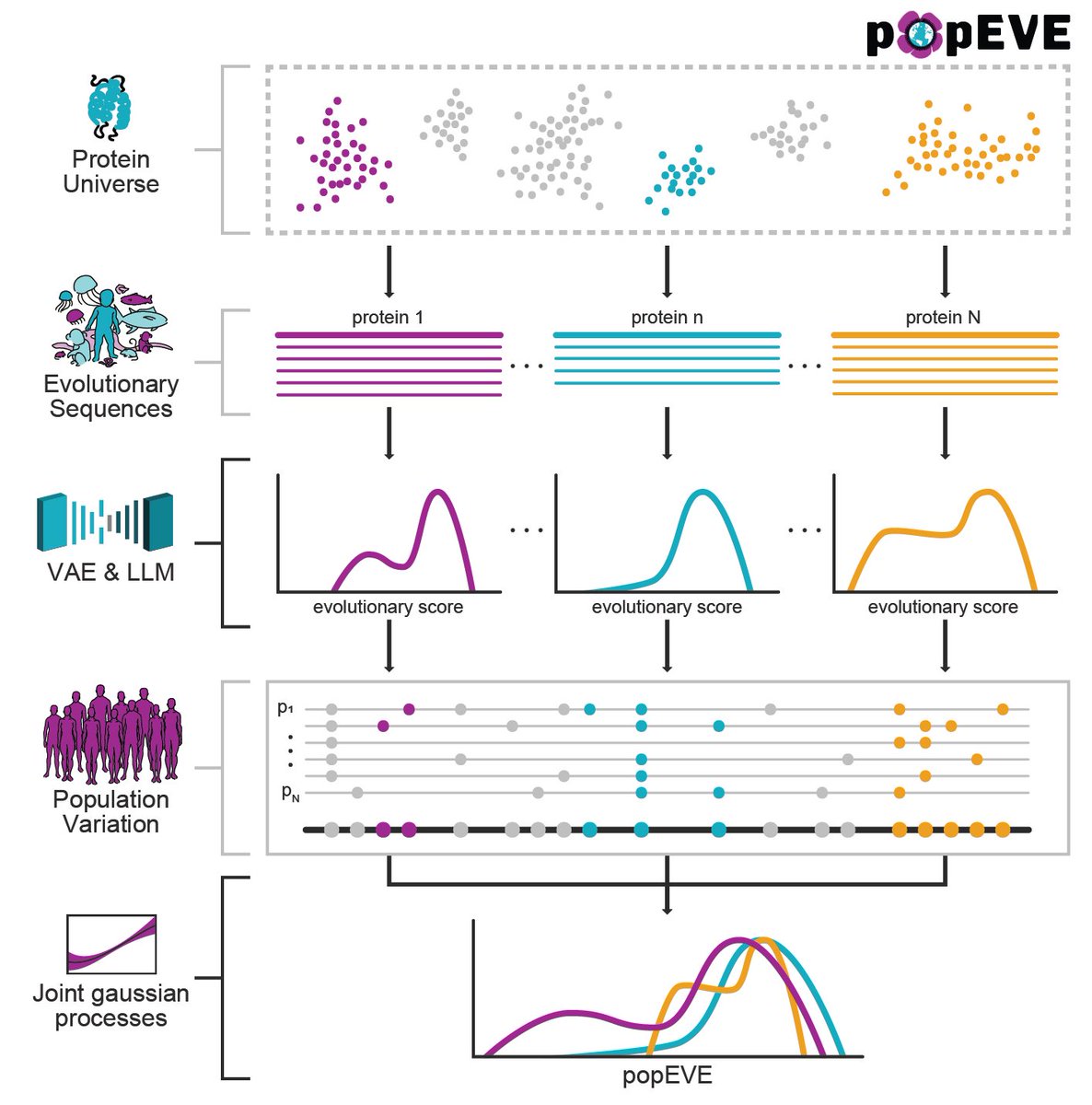
The February issue, with a focus on protein engineering, is live nature.com/nbt/volumes/42… Our cover shows the three data types key to machine learning for functional protein design: structure, sequence and labels, from a Review by Notin et al. go.nature.com/49EPi9x





We're looking forward to presenting preclinical data describing the application of our IMPACT platform at PEGS Boston 2023 #PEGS23 next week, the world’s largest gathering of protein engineering and biotherapeutics experts. #WeAreSesimic seismictx.com/seismic-therap…