abiha retweetledi
abiha
289 posts

abiha
@abihaa__
scientist in the making 🧬🖤 I believe in the power of words. so I write them, read them, and hand letter them 📝📚🎨
Dar es Salaam, Tanzania Katılım Ocak 2018
255 Takip Edilen133 Takipçiler
abiha retweetledi
abiha retweetledi

Today in @Nature a synthesis of the epidemiology of child #malnutrition in the Global South published in 3 papers led by our team. Below, I’ll highlight a few of the results #epitwitter #Medtwitter #tweetorial 🧵 (1/17)
English
abiha retweetledi
abiha retweetledi

Research sheds light on the molecular mechanisms behind CD31 signalling that leads neutrophils to inflamed sites. elifesciences.org/articles/84752…
English
abiha retweetledi

Enamel is tough enough to resist dents, yet elastic enough not to crack during decades of jaw smashing.
It’s so incredible that scientists haven’t created a substitute that can match it—until last year. #ScienceMagArchives scim.ag/3Q8
English
abiha retweetledi
abiha retweetledi
abiha retweetledi

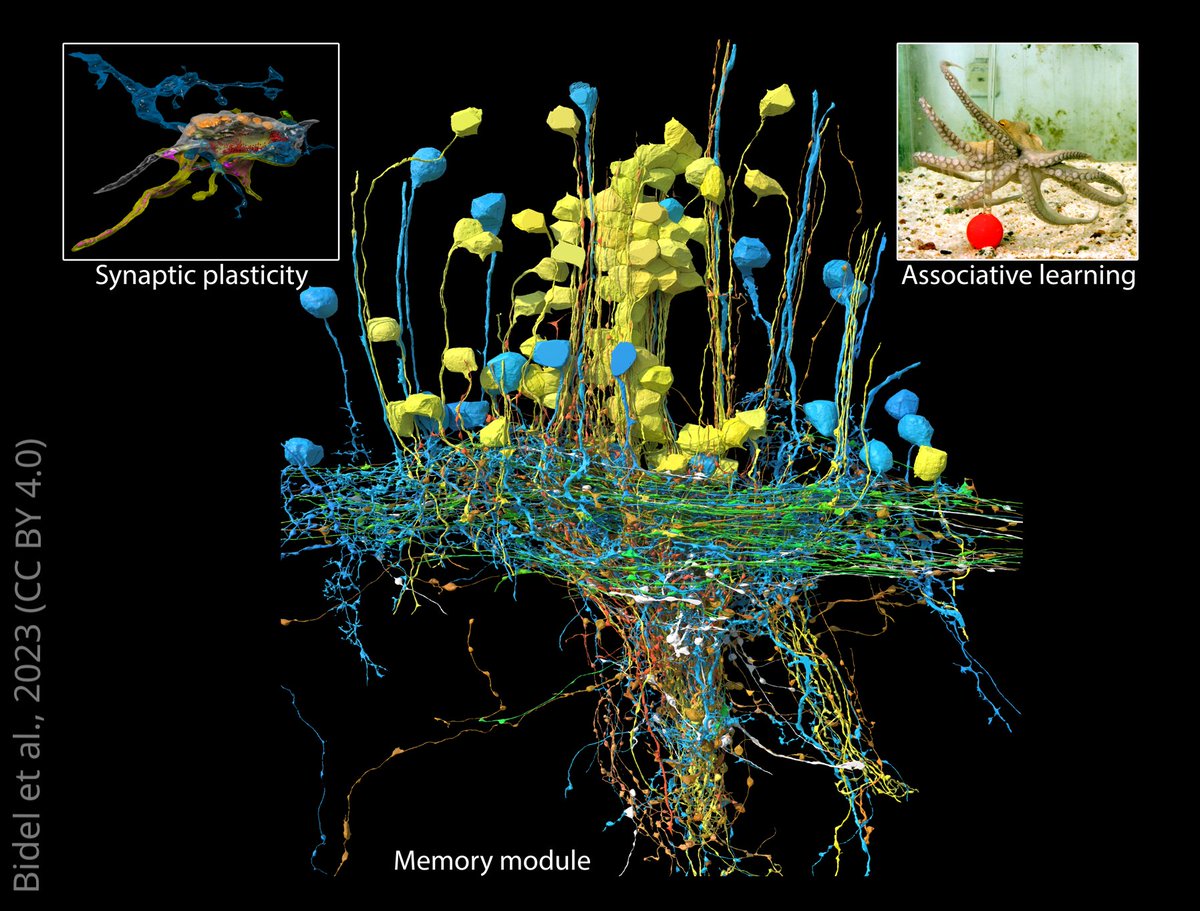
For the first time, scientists have analysed the connectome for part of the brain in the common octopus that’s involved in acquiring long-term memories. elifesciences.org/articles/84257…
English
abiha retweetledi

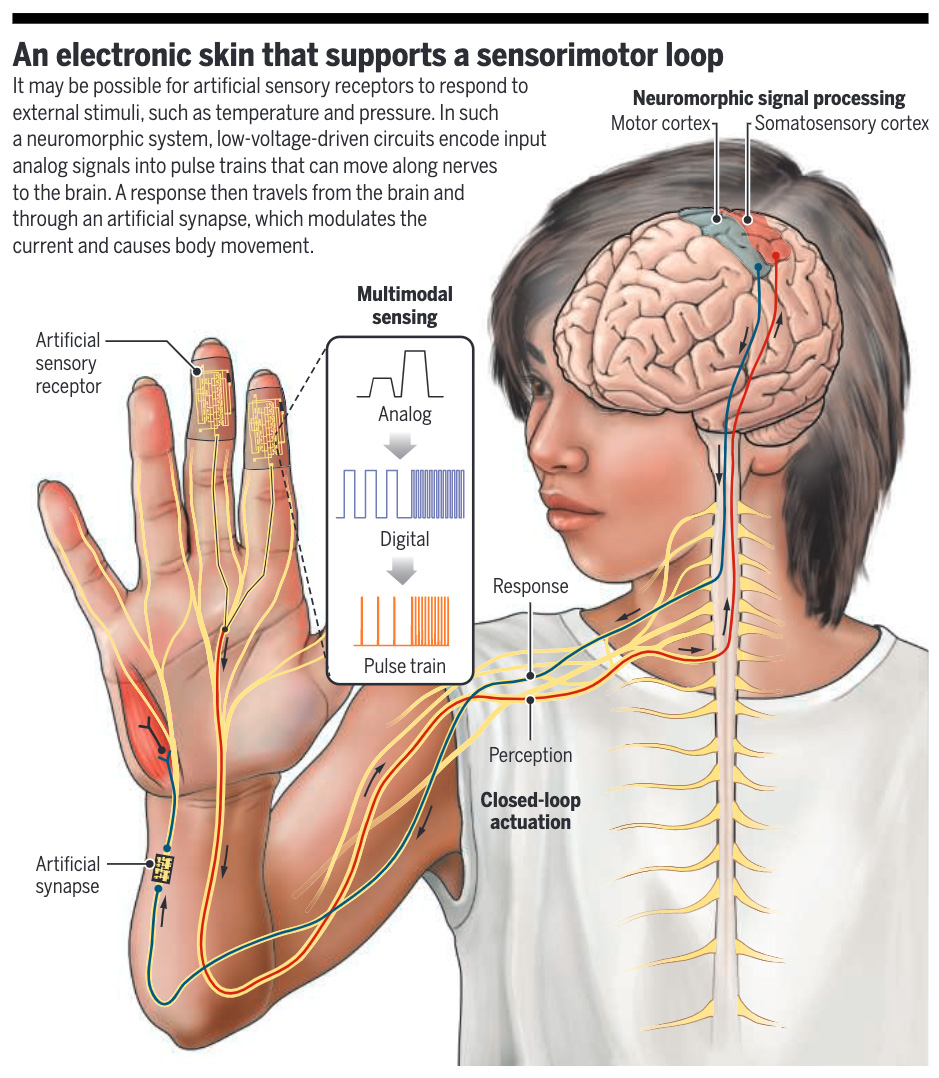
Researchers have developed and tested in rats a neuromorphic electronic skin system that simultaneously emulates closed-loop sensory encoding and the mechanical softness of natural skin.
Learn more in a recent #SciencePerspective: scim.ag/3yE
English
abiha retweetledi

In this study, AI was more accurate than two thirds of radiologists, yet when radiologists had AI help their diagnoses did not improve. Why?
Humans ignored the AI’s advice when it conflicted with their views. A big barrier to future human-AI collaboration blueprintcdn.com/wp-content/upl…


English
abiha retweetledi
abiha retweetledi

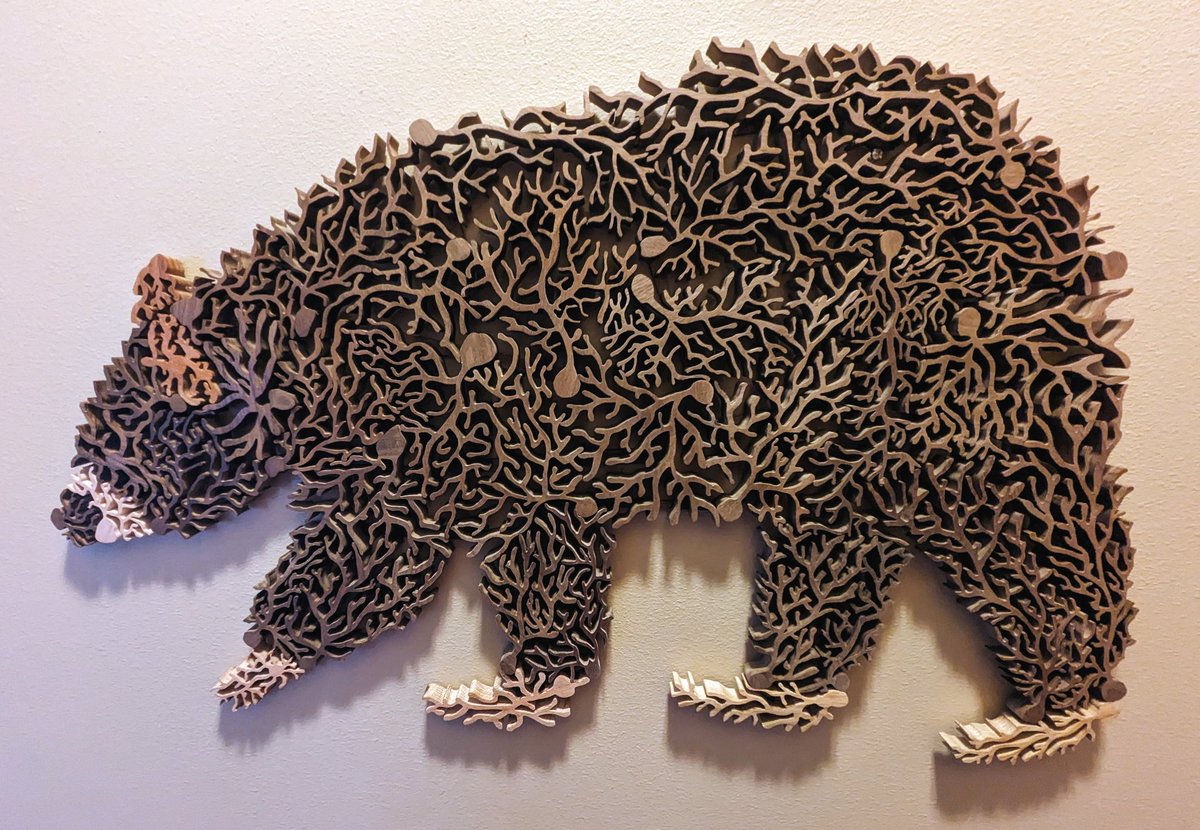
Just finished my most ambitious project yet: "the purkinje bear" 🐻. If you look closer, you will notice that this bear is made of 37 #brain cells, each manually cut with a scroll saw. Walnut, oak, sapele and maple woods. #neuroscience #animal #art

Sioux Falls, SD 🇺🇸 English
abiha retweetledi

Day 8 of great synthetic biology papers.
“Idempotent Vector Design for Standard Assembly of Biobricks” (2003).
Genetic circuits rarely work on first try. It's hard to copy someone else’s work.
So this paper standardized DNA assembly, much like screws were standardized in 1864.
*****
In the 1850s, if you wanted to build a house, you might go chop down some wood and then fasten together the logs using wonky-looking screws, which have been around since at least 400 B.C. but were not standardized until 1864.
A man named William Sellers was responsible. He gave a speech at the Franklin Institute in Philadelphia, proposing to standardize the screw; to make each one, of varying sizes, have a consistent “pitch, diameter, and form of screw threads.”
Up to this point in history, each screw had a different shape. Each workshop made them in a different way.
But, after Sellers' speech, an entire industry quickly formed around the new standards. Soon, all the different workshops made screws in the same way. Suddenly, a blueprint for a bridge in San Francisco could be duplicated in Tokyo or Sydney. Engineering became standardized, consistent, and replicable.
So why should biology be any different?
After all, synthetic biologists spend lots of time building genetic circuits from small, modular sequences. A gene in bacteria is made from a promoter, coding sequence, ribosome binding site, and terminator.
Each of these parts can be isolated from diverse organisms, placed into a cell, and they just...work.
But it wasn't until 1996 that a group of researchers at the University of Illinois at Chicago first proposed consistent standards for assembling DNA in biology! They called their approach, published in PNAS, NOMAD. It stands for Nucleic acid Ordered Module Assembly with Directionality.
“Use of NOMAD ensures that the production of experimental constructs is no longer the rate-limiting step in applications that require combinatorial rearrangement of DNA fragments,” they wrote. (pnas.org/doi/abs/10.107…)
The paper is very impressive. It lays out a whole system to stitch together promoters and other “pieces” into synthetic genes. But nobody really seems to have given a damn, until seven years later, when Tom Knight (who ran a biology lab at the MIT AI Laboratory, and who had previously designed and implemented the MIT Lisp Machine processor and also “supervised the construction of the first PDP-10 ARPANET interfaces with Bob Metcalfe) devised a similar standardization protocol, called Biobricks. (en.wikipedia.org/wiki/Tom_Knigh…)
The basic idea of Biobricks is to store each individual DNA “part” — a promoter, coding sequence, terminator, whatever — on a circular loop of DNA, called a plasmid. Each part is then flanked with short DNA fragments, called restriction sites, that are recognized and cleaved by specific enzymes. When these sites are cut, they leave behind little overhangs that can be stitched together with other DNA parts.
It’s a really clever approach, and useful. This paper, much like Sellers' speech for screws, formed the entire basis by which we now clone and assemble DNA in synthetic biology.
h/t @jrkelly
Paper: dspace.mit.edu/handle/1721.1/…

English
abiha retweetledi
abiha retweetledi
abiha retweetledi
abiha retweetledi
abiha retweetledi

@andrewmccalip Wait…are you trying to implement the paper rn?
English












