Brandon Logeman
295 posts


Brandon Logeman
@brandon_logeman
BRAIN Initiative K99/R00 Awardee || Studying hormonal control of neuronal circuits at the single cell level || 🐷 former pig farmer 🐷 #FirstGenCollege










I’m looking for examples where a gene is normally expressed in cell type A but a GWAS/QTL hit contains a variant that causes ectopic expression in cell type B. All examples welcome, plz RT! @jkpritch @doctorveera @SashaGusevPosts @dgmacarthur @tuuliel @anshulkundaje




The most persistent problems in science trace directly to the fact that many of its practitioners hold on to banal utopian visions of how it should operate and actively choose not to engage with the reality of how it actually works and how it fits into society at large.








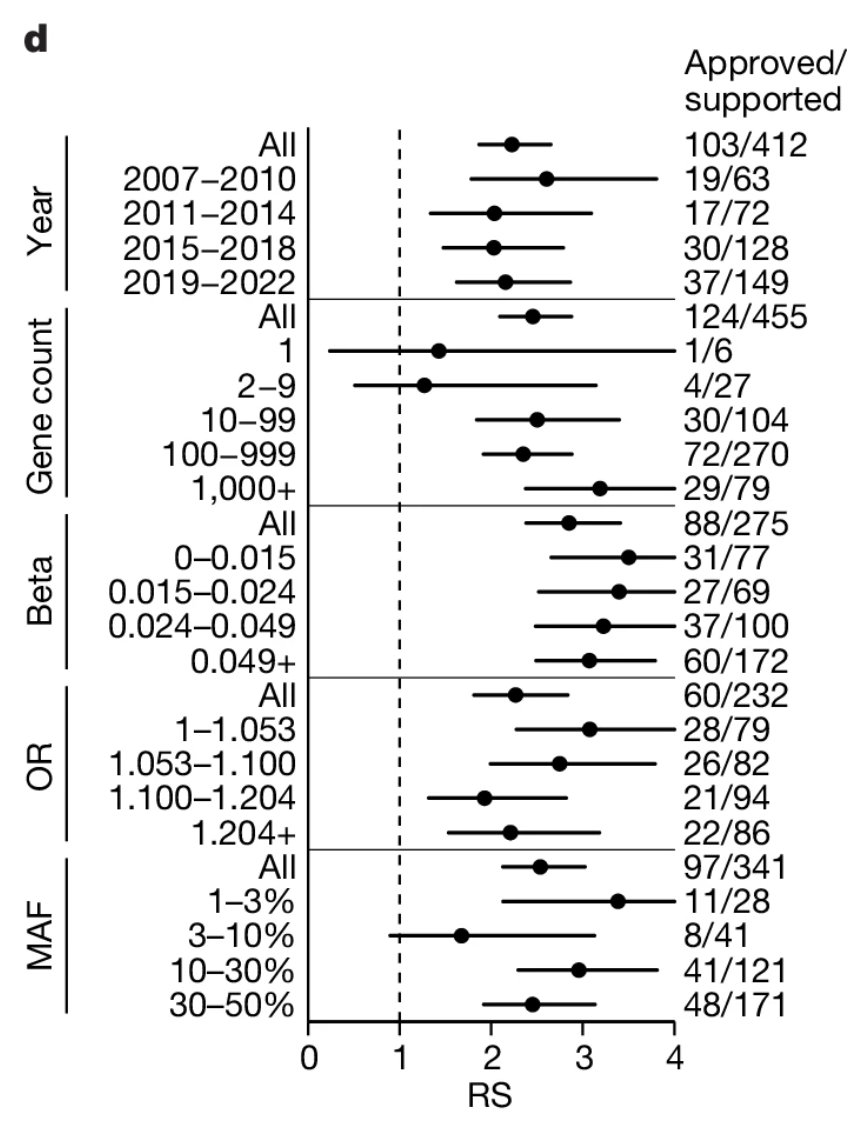
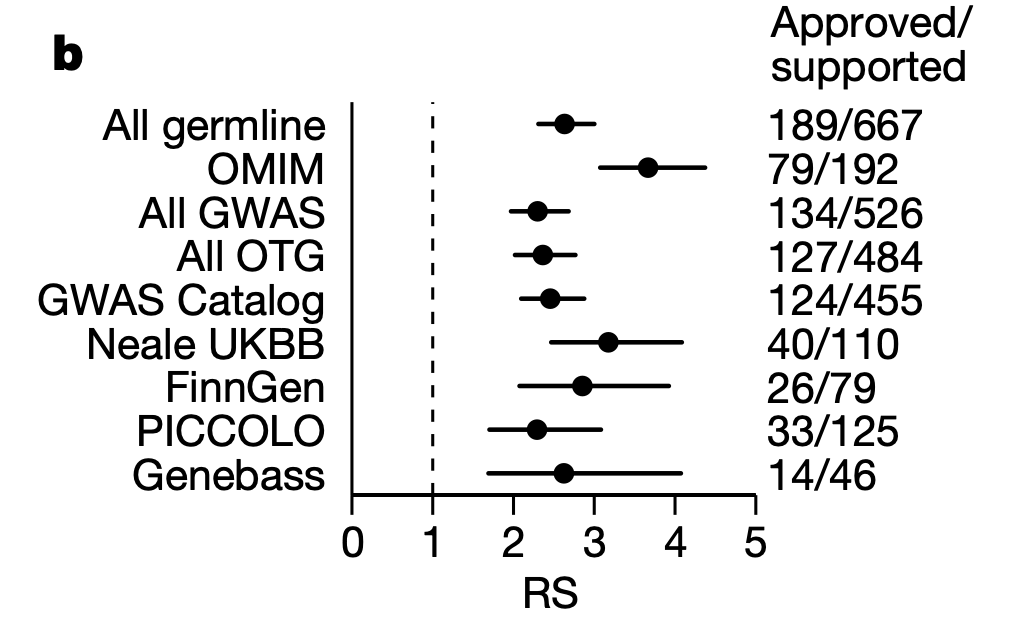


This recent GWAS preprint is the first to explore genetic influences on the age at onset of walking 🚶♀️🚶, a milestone that may indicate broader (neuro)developmental delays. Find out more⤵️ medrxiv.org/content/10.110…