Brad Chapman
1.6K posts

Brad Chapman
@chapmanb
Biologist and programmer @chapmanb.bsky.social @[email protected]


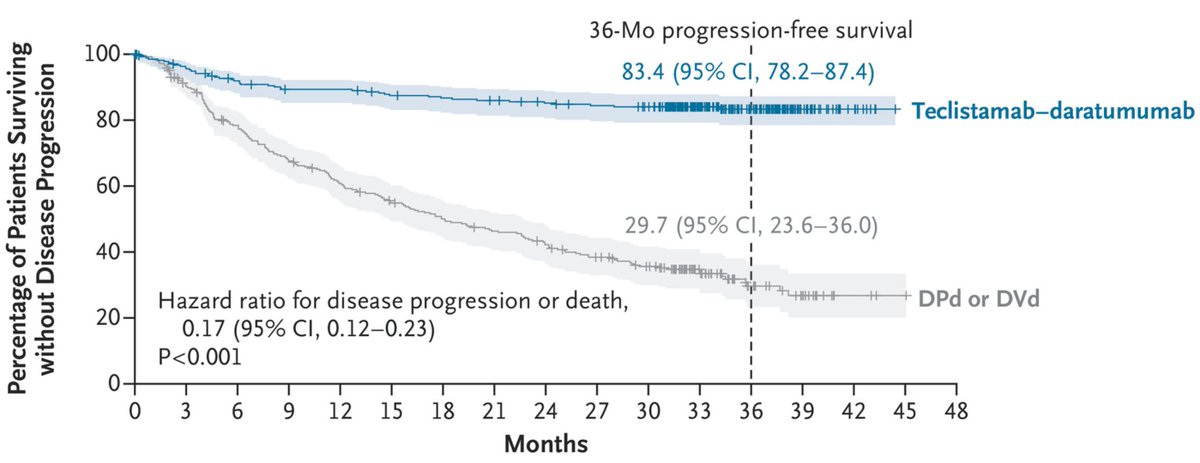
Encouraging news for patients living with multiple myeloma. The FDA has expanded the use of a combination treatment for multiple myeloma, allowing it to be used earlier when the disease returns or starts to worsen. “This approval is due to years of scientific progress and will be a meaningful improvement for many patients,” says Dr. Gruenbaum. Learn more: bloodcancerunited.org/resources/news…

We used AI to predict the failure of a Phase 3 trial before the results were announced. Today, we're publishing 10 more predictions for the future. Thread 🧵

Excited to share our new Nature Methods paper on longcallR, a Rust tool for long-read RNA-seq that jointly performs SNP calling and phasing, and enables haplotype-specific junction analysis. Thanks to @lh3lh3 for the support and guidance. Paper: nature.com/articles/s4159…

Myloasm, our long-read metagenome assembler, is now published! w/ Max Marin & @lh3lh3 Very rewarding after > a year of development and countless hours thinking about assembly. Thanks to beta testers, Li lab, and reviewers for helpful feedback. Link: rdcu.be/famFj


















A few years ago, designing an antibody on the computer was extremely difficult. Today, there are several open-source tools which allow anyone to design antibodies from home. Out today: A step-by-step guide to antibody design. By @btnaughton.