Gerald Tiu retweetledi

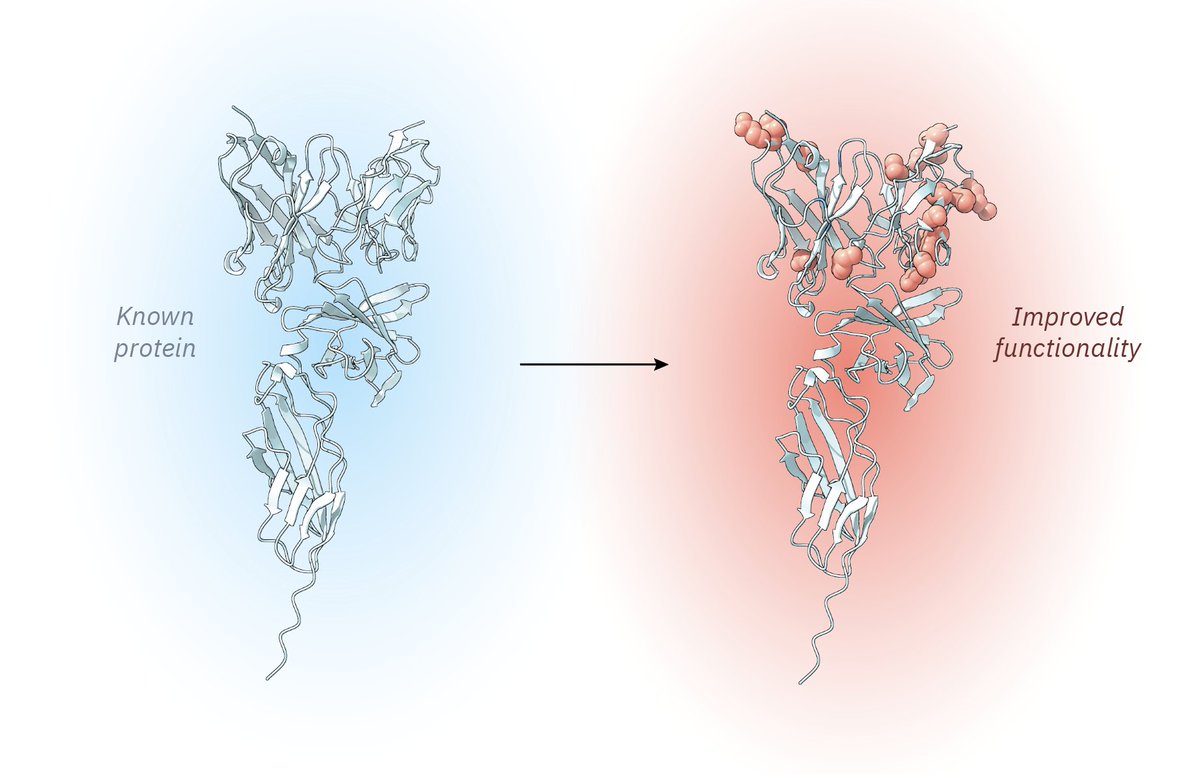
The hardest part of protein engineering isn't just finding good mutations – it’s deciphering which ones combine synergistically.
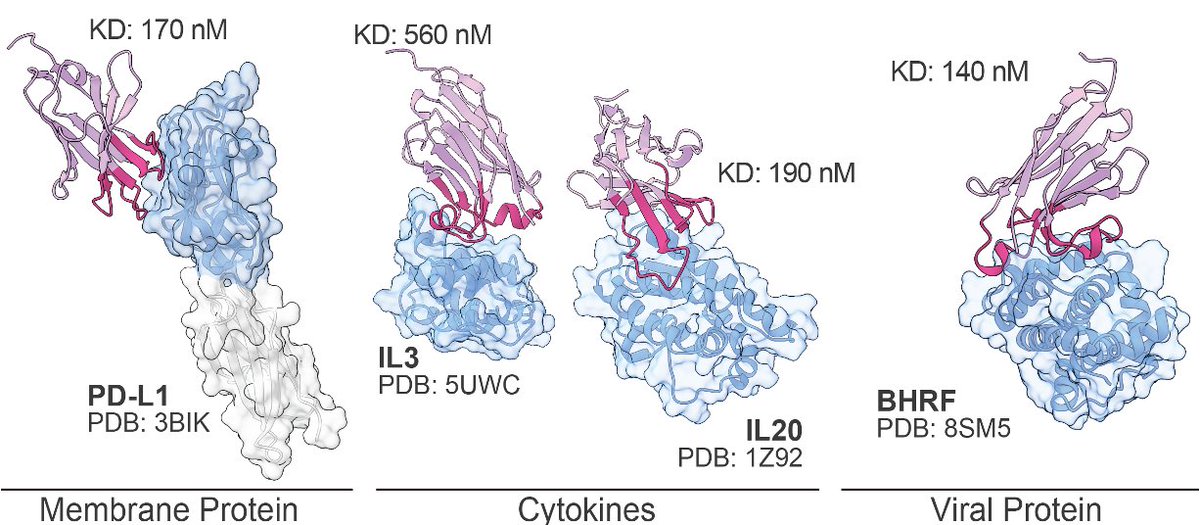
Today in @ScienceMagazine, we present MULTI-evolve, a framework for rapid multi-mutant protein engineering, validated across three diverse proteins.
English