Jeff Nirschl
773 posts

Jeff Nirschl
@jnirsch
M.D.–Ph.D. interested in computational image analysis, digital pathology, and neuropathology. Personal account: all opinions are my own.


🚀 Introducing iSight: toward expert–AI collaboration for #immunohistochemistry (IHC) assessment Preprint: arxiv.org/abs/2602.04063 🚀 What we built: 🖼️ HPA10M, the world’s largest open-access IHC dataset: huggingface.co/datasets/nirsc… • 10,495,672 IHC images (Over 10M!) • 17,000+ protein markers • 45 normal tissues + 20 cancer types • Fully curated, standardized, and publicly available on @huggingface. We trained iSight, a multi-task AI system for automated IHC assessment • Jointly predicts staining location, intensity, and quantity • Outperforms fine-tuned pathology foundation models • Produces well-calibrated, clinically interpretable outputs @PennPathLabMed #DigitalPathology #AIinHealthcare #Immunohistochemistry #HumanAI #Pathology

Skin cancer diagnosis isn’t just about seeing. It’s about knowing what you’re seeing. A new AI system improves melanoma diagnosis by combining pathology images with expert knowledge, not just pixels. READ HERE: nature.com/articles/s4174…






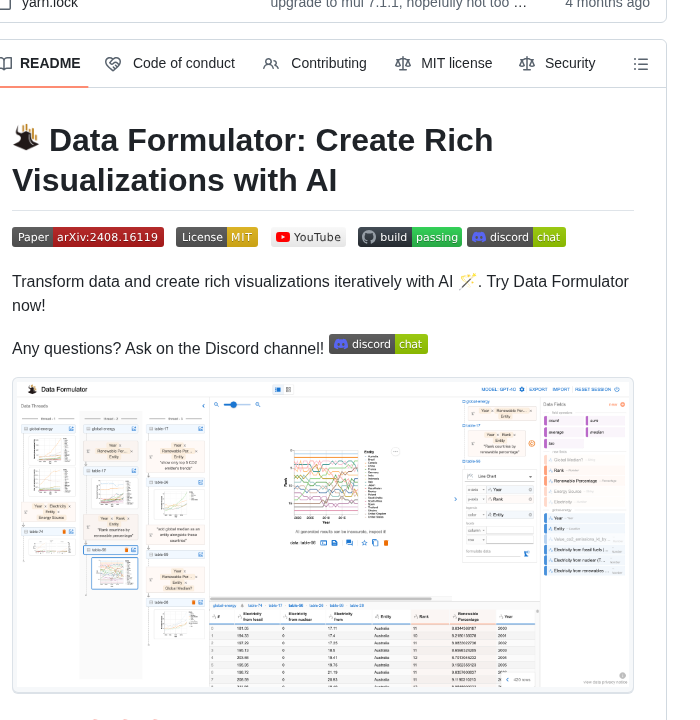
Recently we released a 3000+ word book chapter written by @KeremTurgutlu, based on @karpathy's marvelous "Let's built the GPT tokenizer" video. It's got pics, links, code, diagrams, … Kerem has now written a detailed walk-through of how he made it: answer.ai/posts/2025-10-…

Wow! TissueLab.org has reached over 1700 users in the first 30 days and almost 500 downloads 🚀🔥! What an impressive achievement by my team. So proud of my students! We will keep shipping the code, fixing the bugs, and improve user experiences. Check out the first co-evolving agentic AI system for medical image analysis! 👉arxiv.org/abs/2509.20279 #TissueLab #AIAgent #AgenticAI #pathology #radiology

BERT is just a Single Text Diffusion Step! (1/n) When I first read about language diffusion models, I was surprised to find that their training objective was just a generalization of masked language modeling (MLM), something we’ve been doing since BERT from 2018. The first thought I had was, “can we finetune a BERT-like model to do text generation?”