Sabitlenmiş Tweet
Mitchell B Slapik
484 posts


Mitchell B Slapik
@mslapik
MD/PhD Candidate 🥼 @McGovernMed | F30/TL1 Fellow 🏥 @NIH | Computational Neuroscience 🧠 @DragoiLab | Jazz Sax 🎷 @TheChirpChirps
Houston, TX Katılım Kasım 2022
961 Takip Edilen3K Takipçiler
Mitchell B Slapik retweetledi

1/ New preprint with @dyamins + team! Ventral visual representations within areas evolve over the course of the response along the same hierarchical complexity axis that distinguishes the visual areas, potentially driven by local recurrence.
biorxiv.org/content/10.648…

English
Mitchell B Slapik retweetledi
Mitchell B Slapik retweetledi
Mitchell B Slapik retweetledi
Mitchell B Slapik retweetledi
Mitchell B Slapik retweetledi
Mitchell B Slapik retweetledi

🧵1/ Our new study on AI and physician reasoning just came out in @ScienceMagazine. As co-senior author, I'm excited about our findings, and I do think AI will reshape medicine. But after seeing some of the discussions, I'm also worried about how our findings may be misinterpreted.

English
Mitchell B Slapik retweetledi

My bet: in the near future, 80%⬆️ of CS research will be done by AI in collaboration with humans. However, today's research ecosystem is still built around the human, not the AI scientist.
For example, the 8-page paper PDF is a lossy compression of months of branching exploration into a linear story, optimized for a human reviewer to skim in 30 minutes. It hides two structural taxes:
📖 Storytelling Tax — failures, rejected hypotheses, and dead ends get stripped. On RE-Bench (24,008 runs, 21 frontier models), failed runs = 90.2% of total compute cost, with a 113× median failed-to-success token ratio. Every lab independently rediscovers the same dead ends.
🔧 Engineering Tax — the gap between reviewer-sufficient prose and agent-sufficient spec. Across 8,921 PaperBench requirements (23 ICML'24 papers), only 45.4% are fully specified in the PDF. The rest is tacit lab knowledge. Tolerable when readers were human. Critical now that agents read, reproduce, and extend.
We propose ARA: the Agent-Native Research Artifact — replace the narrative PDF with an agent-executable package, in 4 layers:
🧠 structured scientific logic
⚙️ executable code w/ full specs
🌳 exploration graph (every failure preserved)
📊 evidence grounding every claim
English
Mitchell B Slapik retweetledi

What if the model didn’t just use a computer, but actually was the computer?
Meta AI introduces "Neural Computer", a model where computation, memory, and I/O are all inside one learned system.
Their early prototype learns from screen recordings of terminals and desktops, and it can already imitate some basic computer behavior like rendering interfaces and responding to clicks or commands.
But it still breaks on slightly harder tasks like reliable reasoning, stable memory, and reusable skills.

English

My new paper on "AI Consciousness".
I argue that to talk about that we need to have *validated* theories of human consciousness (and we're very far from that)
Accepted at AAAI Symposium 2026 on Machine Consciousness.
Feedback and discussion welcome.
Lossfunk@lossfunk
🚨 New Paper Can AI models be conscious? We argue that answering this question requires us to have a validated theory of human consciousness first and without that, the concept “ai consciousness” is not well grounded. Accepted at AAAI Symposium 2026 lossfunk.com/papers/ai-cons… 🧵
English
Mitchell B Slapik retweetledi

🚨 New Paper
Can AI models be conscious?
We argue that answering this question requires us to have a validated theory of human consciousness first and without that, the concept “ai consciousness” is not well grounded.
Accepted at AAAI Symposium 2026
lossfunk.com/papers/ai-cons…
🧵

English
Mitchell B Slapik retweetledi

Across versions of ChatGPT, responses to psychotic prompts were frequently inappropriate or partially appropriate, raising safety concerns for users at risk for #psychosis.
ja.ma/4mb41Ae

English
Mitchell B Slapik retweetledi

Psychiatry is at a crossroads.
What if much of what we diagnose as disorder is actually a rational response to an unstable world shaped by climate change, conflict, and uncertainty?
Our new editorial 👇
globalpsychiatryarchives.com/index.php/gpa/…

English
Mitchell B Slapik retweetledi

Postpartum depression is often missed, & not all cases look the same.
A new study in @BMJMentalHealth shows AI can help: by analysing clinical notes, researchers identified 30% more PPD cases beyond diagnosis codes alone, revealing distinct subtypes with different needs.
Earlier detection = better, more personalised care
Link: mentalhealth.bmj.com/content/29/1/e…
Authors:
Prakash Adekkanattu, Veer Vekaria, Yiye Zhang, Braja Gopal Patra, Priscilla Liang, Marianne Sharko, Natalie Benda, Meghan Reading Turchioe, Andrea Temkin- Yu, Alison Hermann, Jyotishman Pathak
@Columbia

English
Mitchell B Slapik retweetledi

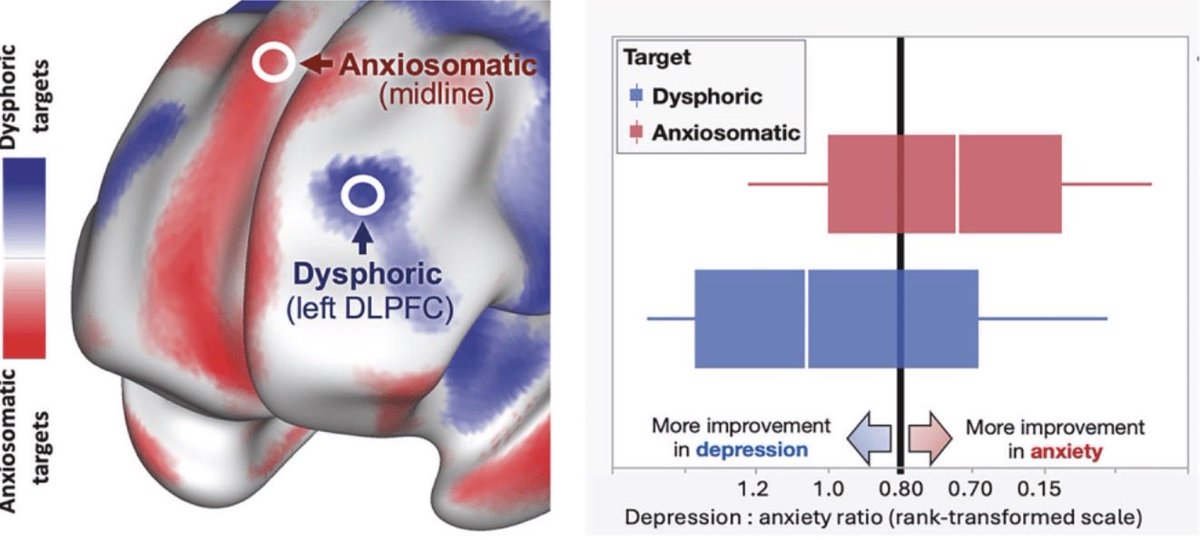
Our latest in @molpsychiatry: In patients with anxiety+depression, targeting a novel “anxiosomatic” circuit in dmPFC outperforms standard dlPFC target for anxiety, and equally effective for depression.
nature.com/articles/s4138…
English
Mitchell B Slapik retweetledi

Among individuals with severe, treatment-resistant #Schizophrenia, #dementia was common and showed a distinct clinical and genetic profile not explained by #Alzheimer disease, cardiovascular risk, or medication effects.
ja.ma/4uWAsGG

English
Mitchell B Slapik retweetledi
Mitchell B Slapik retweetledi
Mitchell B Slapik retweetledi

The missing half of the neural network–brain comparison
For a decade, the standard benchmark for artificial neural networks as models of the brain has been forward predictivity: learn a linear mapping from model activations to neural recordings and measure explained variance. Top models of the macaque inferior temporal (IT) cortex—central to object recognition—have plateaued near 50% regardless of architecture.
Muzellec and Kar argue this plateau hides something important. Two models can score identically on forward predictivity while relying on fundamentally different internal strategies. One may have many units tightly coupled to IT responses; the other may reach the same score with a smaller aligned subset while carrying a large pool of biologically inaccessible dimensions.
To expose this, they introduce reverse predictivity: instead of asking how well model features predict neurons, they ask how well IT neurons predict individual model units. A truly brain-like model should be bidirectionally predictable—just as two monkeys' IT populations predict each other symmetrically, which the authors confirm as their empirical baseline.
Across 39 architectures—CNNs, transformers, self-supervised and robust models—reverse predictivity is consistently lower than forward predictivity and the two metrics are uncorrelated. Strikingly, higher ImageNet accuracy predicts lower reverse predictivity. Adversarial training helps; higher dimensionality hurts. The "common" units identified this way predict primate behavior more consistently across species and models than the "unique" ones inaccessible from neural activity.
For AI in drug discovery, neurotechnology, or computational biology, this has a direct implication: forward accuracy alone does not guarantee that a model's internal representations are embedded in the biological system it claims to describe. When those representations guide mechanistic interpretations or experimental decisions, the mismatch can mislead.
Paper: Muzellec et al., Nature Machine Intelligence (2026) | nature.com/articles/s4225…

English
Mitchell B Slapik retweetledi

"Before we name, we touch. We propose that the roots of language lie not in abstract, amodal symbols but in early bodily experience." Our Opinion paper is out in TICS. Congrats Luca Sergey Rinaldi !!
sciencedirect.com/science/articl…




English










