Sabitlenmiş Tweet

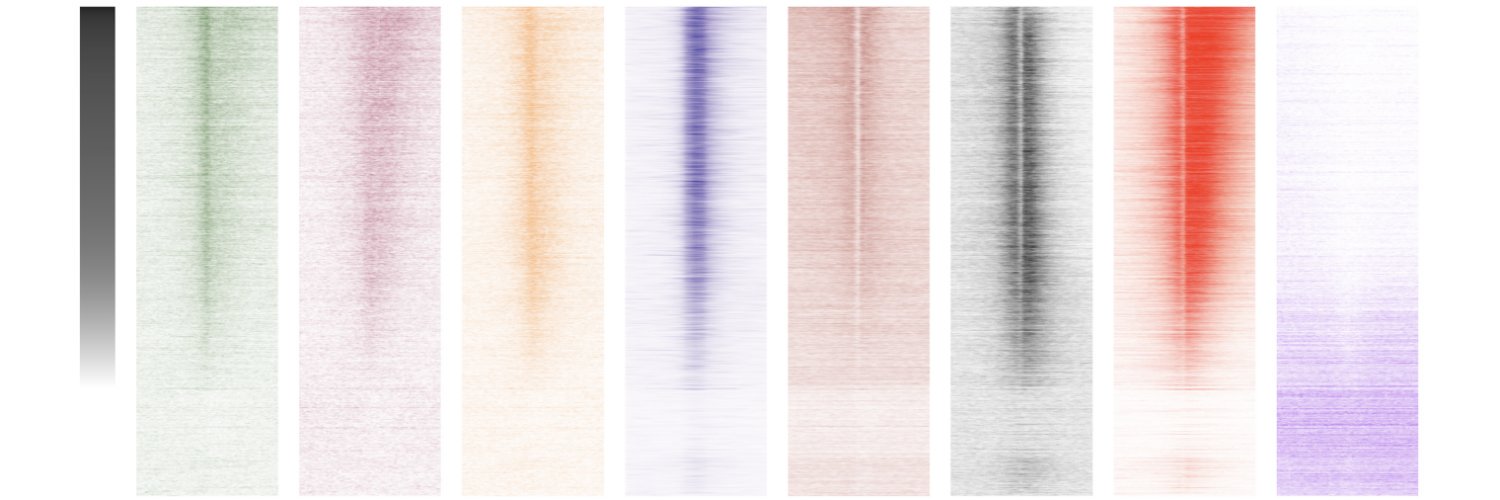
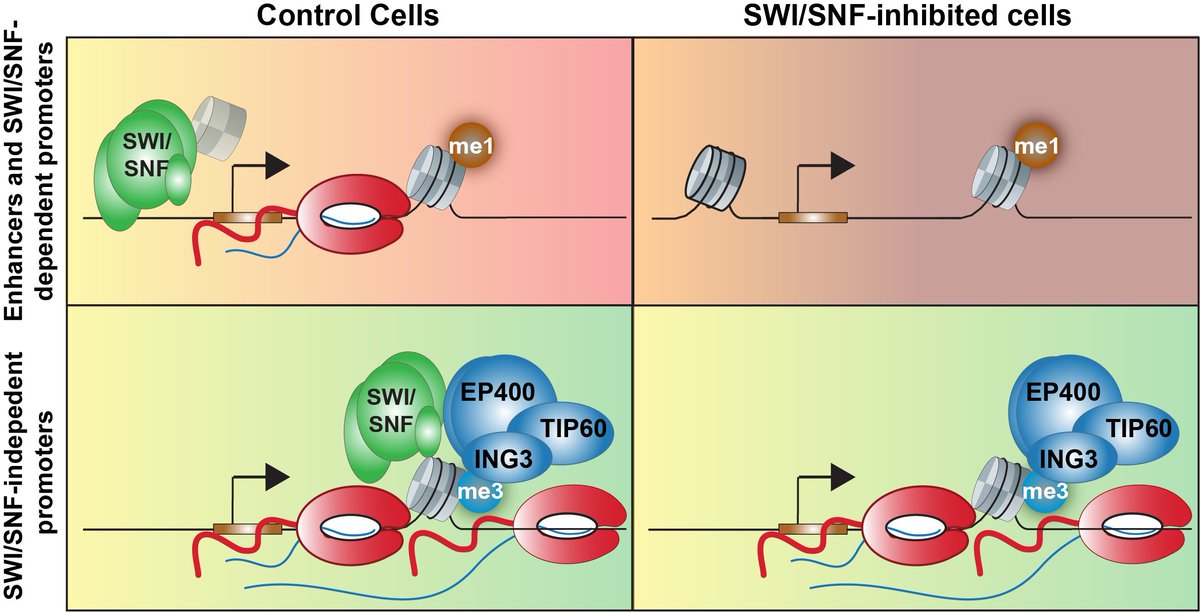
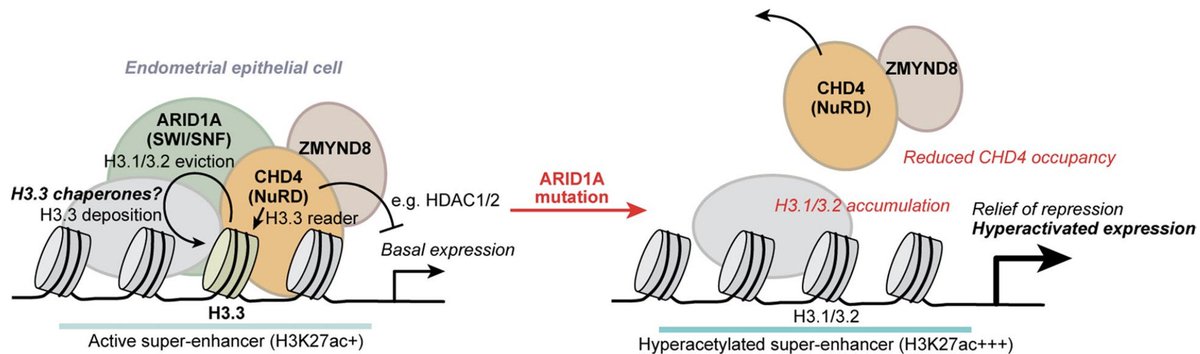
A chromatin mechanism hiding in plain sight. We found that a fundamental function of mammalian ARID1A-SWI/SNF chromatin remodeling is to facilitate H3.3 variant histone exchange, likely by ejecting canonical H3.1/3.2.
bmcbiol.biomedcentral.com/articles/10.11…
English