Rob Douglas
37.6K posts


Rob Douglas
@RobDouglas
ID Theft & Fraud Prevention Consultant ~ Aspiring Gym Rat ~ Fmr Private Detective ~ Fmr Political Commentator ~ Student of Stoicism ~ Disciple of Nature

The human proteome just expanded by thousands of proteins—and most of them are sized perfectly for peptide therapeutics. A new Nature study from the TransCODE Consortium analyzed 95,520 proteomics experiments and found that approximately 25% of 7,264 non-canonical open reading frames encode detectable microproteins in human cells. These sequences produce peptides ranging from 8 to 100 amino acids. That's the therapeutic window where synthetic peptides can be manufactured, modified for stability, and delivered as drugs. Peptide therapeutics have been constrained by the need to target biologically active sequences that are small enough to synthesize but specific enough to produce therapeutic effects. The challenge has been identifying which endogenous peptides perform essential cellular functions and which can be modified into pharmacologically stable molecules. This study systematically maps thousands of naturally occurring peptides that cells translate and use across tissues and disease states. Many were detected in the HLA PeptideAtlas—240 million mass spectrometry spectra from immunopeptidomics datasets showing which peptides cells present on their surface. That's direct evidence of biological relevance. Cells don't randomly present peptides on HLA molecules. Presentation indicates processing, translation, and integration into cellular signaling or immune surveillance. The therapeutic implications operate across multiple modalities. First: peptide replacement therapy. If microproteins perform essential cellular functions but decline with age or disease, synthetic versions could restore activity. This mirrors the logic behind hormone replacement—when endogenous production drops, exogenous supplementation compensates. The consortium demonstrated that one peptidein from the OLMALINC long non-coding RNA produces a pan-essential cellular phenotype. CRISPR screens showed its loss disrupts fundamental processes. That's a candidate for peptide replacement if expression declines in specific tissues or aging contexts. Other microproteins may regulate mitochondrial function, proteostasis, or cellular senescence. If age-related decline in these peptides contributes to metabolic dysfunction, peptide-based interventions could target those pathways directly rather than modulating upstream regulatory machinery. Second: peptide antagonists. Some microproteins may drive pathological processes—inflammatory signaling, oncogenic pathways, or maladaptive stress responses. Designing antagonist peptides that block microprotein activity creates therapeutic options for conditions where microprotein overexpression or dysregulation contributes to disease. The study found cancer-specific microproteins expressed in malignant cells but not normal tissues. These represent targets for peptide-based inhibitors that disrupt cancer cell signaling without affecting healthy cells. Because these sequences aren't part of the canonical proteome, conventional small molecule screens wouldn't have identified them. Third: peptide vaccines. The HLA PeptideAtlas detected peptides from 1,785 ncORFs presented on cell surfaces. Cancer cells presenting unique microprotein-derived peptides expose targetable antigens for therapeutic vaccines. This approach already exists with neoantigens—tumor-specific mutations that generate novel peptides recognized by T cells. Microprotein-derived peptides expand that target space. They're not mutations—they're translation products from sequences that normal cells suppress but cancer cells express. Peptide vaccines could train the immune system to recognize and eliminate cells presenting these cryptic antigens. Because microproteins are often cancer-restricted, this strategy may produce stronger anti-tumor responses with fewer autoimmune risks than vaccines targeting overexpressed canonical proteins. Fourth: cell-penetrating peptides and delivery vehicles. Some microproteins may function as endogenous cell-penetrating sequences—naturally occurring peptides that cross membranes or localize to specific organelles. Identifying these sequences could improve drug delivery technology. Current peptide therapeutics face bioavailability challenges. Oral delivery is difficult due to enzymatic degradation. Systemic delivery requires modifications to extend half-life and prevent renal clearance. Intracellular targeting remains complex because most peptides don't efficiently cross lipid bilayers. If evolution has already produced microproteins with membrane-crossing or organelle-targeting capabilities, those sequences could be incorporated into therapeutic peptides to improve cellular uptake and subcellular localization. Fifth: synthetic biology and designed peptides. The study provides a catalog of naturally occurring bioactive peptides that cells translate and tolerate. That catalog becomes a training set for designing synthetic peptides with desired pharmacological properties. Machine learning models trained on microprotein sequences—combined with data on their tissue expression, HLA presentation, and evolutionary constraint—could predict which synthetic peptide sequences will be stable, non-immunogenic, and biologically active. This accelerates peptide drug development by narrowing the design space. Rather than screening random sequences, developers can modify known functional microproteins or generate synthetic analogs based on evolutionary patterns. The manufacturing advantage: peptide synthesis is straightforward. Unlike biologics requiring expression systems and purification pipelines, peptides can be chemically synthesized at scale. Modifications to enhance stability—D-amino acids, cyclization, lipidation—are well-established. The pharmacokinetic challenge has been specificity and half-life. Endogenous microproteins solve the specificity problem—they're already performing targeted cellular functions. Engineering modifications to extend circulation time becomes the primary optimization. The evolutionary analysis supports therapeutic viability. The consortium developed ORF relative branch length (ORBL) to measure selective constraint on microproteins. Sequences under purifying selection across mammalian evolution are preserved because they perform functions that natural selection maintains. That's evidence these peptides matter biologically. Therapeutic interventions modulating microprotein activity aren't targeting random noise—they're engaging functional molecules shaped by millions of years of selection pressure. The annotation framework enables systematic peptide therapeutic development. By formalizing peptideins as a recognized classification in GENCODE and PeptideAtlas, the consortium creates searchable databases where researchers can identify microproteins relevant to specific diseases, tissues, or cellular processes. Pharma companies developing peptide therapeutics can now query: which microproteins are dysregulated in this disease? Which are cancer-specific? Which show tissue-restricted expression? Which are presented on HLA in patient samples? Those queries weren't possible when microproteins remained unannotated. Now they're part of the reference proteome. The clinical development timeline depends on functional validation. Demonstrating that a microprotein performs a therapeutically relevant function—and that modulating it produces measurable clinical benefits—requires the same rigor as conventional drug development. But the discovery phase just accelerated. Instead of screening millions of synthetic peptides for activity, researchers can start with endogenous sequences that cells already use. The study detected 183 ncORFs with high-confidence peptide evidence in conventional samples and 1,785 in HLA immunopeptidomics. That's thousands of potential therapeutic leads—some for replacement, some for antagonism, some for immune targeting. The immediate research agenda involves characterizing which microproteins show disease-specific expression patterns, which can be chemically synthesized with therapeutic stability, and which produce pharmacological effects when administered exogenously. The decisions about which microproteins to develop as therapeutics will depend on target validation showing that the peptide performs a function relevant to human disease and that synthetic versions can recapitulate or block that function. Peptide therapeutics have been limited by the need to find biologically relevant short sequences. This study just mapped thousands of them.




@DoNotComply23 I can comment and like but not post!



We have another kit with yet another new label that testing has shown tirzepatide not Reta. Someone called Yula is the vendor.




If you’re trying to build your physique, lose fat, or simply improve your cognitive and physical performance, not to mention increase your work capacity, and you’re doing anything remotely close to low-carb, you are sabotaging yourself. Intermittent fasting is absolute garbage. You’re just using the clock to reduce your calorie intake and simultaneously torpedoing your metabolic health and performance. Keto, outside of those who’ve made the prior commitment to becoming metabolically fluid, is absolute garbage. And anything low-carb outside of those with severe metabolic disorders or insulin resistance is absolute garbage as well. Carbohydrates are the cleanest fuel you can run on. Carbs refill muscle glycogen, which is what actually powers a hard training session, and they keep your central nervous system fed so your output in the gym and your output at work both stay sharp. Fat and ketones can keep you alive. Carbs let you perform. Eat more carbohydrates. Time your macros properly. But don’t deprive yourself of the finest form of energy you can feed yourself with.






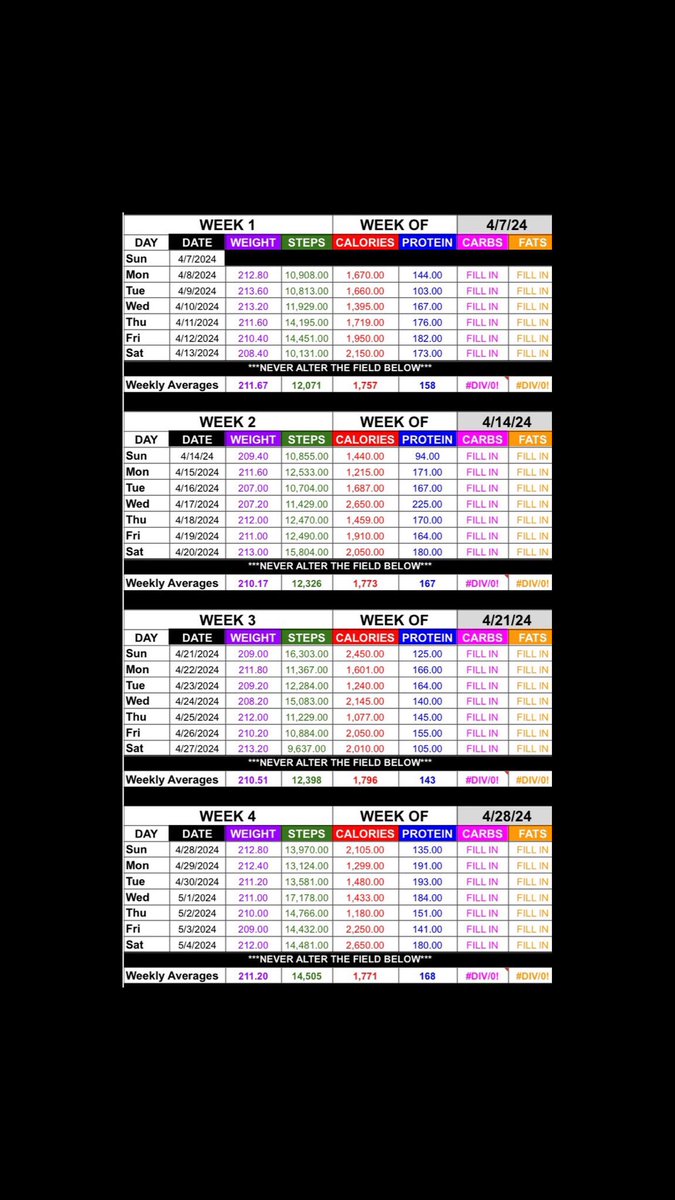

29 out of 30 obese men claimed they ate ~2500 kcal while actually eating ~4000 kcal a day. They were off 37%, ~1500 kcal. Enough to lose 3 lbs per week. Even when they knew they were being watched, they still underreported almost 500 kcal per day. The first step to losing fat is facing the truth about your intake and tracking your calories. If you can't do that, everything else fails.