Artem Kushner
944 posts



@GabriellaG439 gotta say thanks for introducing me to the Dijkstra archive, been reading the EWDs on and off toda.
English

New blog post: "A sufficiently detailed spec is code"
I wrote this because I was tired of people claiming that the future of agentic coding is thoughtful specification work. As I show in the post, the reality devolves into slop pseudocode
haskellforall.com/2026/03/a-suff…
English

this pairs well with that recent epiplexity paper
gabby@GabriellaG439
New blog post: "A sufficiently detailed spec is code" I wrote this because I was tired of people claiming that the future of agentic coding is thoughtful specification work. As I show in the post, the reality devolves into slop pseudocode haskellforall.com/2026/03/a-suff…
English
Artem Kushner retweetledi

📢📢 Proteina-Complexa 📢📢
Atomistic Binder Design with Generative Pretraining and Test-Time Compute + Experimental Validation at Scale
⭐️ Project page (research.nvidia.com/labs/genair/pr…) for:
📜 Method paper (ICLR 2026 Oral)
🧬 Wet lab paper
🛠️ Code & models
📁 Data
🧵 Thread
(1/n)
English
Artem Kushner retweetledi

Radial.org is live! A new organization at @AsteraInstitute
bringing together structural biologists, engineers, and ML scientists to redesign how we do science.
@statnews has the story: statnews.com/2026/03/11/rad…

English
Artem Kushner retweetledi

If you are working with the ribosome and are interested in the shape of its exit tunnel (for simulation, npet dynamics, etc.) here is a tool to get an Angstrom-fine mesh of it in under a minute.
github.com/bioshape-analy…
English
Artem Kushner retweetledi

You like discrete diffusion, but it's too slow? 🥀
You like test-time inference, but it's for continuous methods? 😩
We fixed it.
Introducing Categorical Flow Maps: continuously sample discrete data in a single step 🚀💫
How? 🧵⬇️
💪 Co-led with @FEijkelboom, @daan_roos_
English
Artem Kushner retweetledi

NEW BLOG!
@RuxandraTeslo, @kroetscha, @vientsek, @NeuroStats, and I have started a joint blog on Clinical Trials Abundance!
We'll aim to publish weekly thoughts, commentary and ideas on how to make clinical trials more efficient & abundant.
Subscribe: clinicaltrialsabundance.blog

English

@AnthropicAI i realize all the big boys are using claude code, but it would be very nice if one could collapse long code snippets in the claude.ai app.
English
Artem Kushner retweetledi

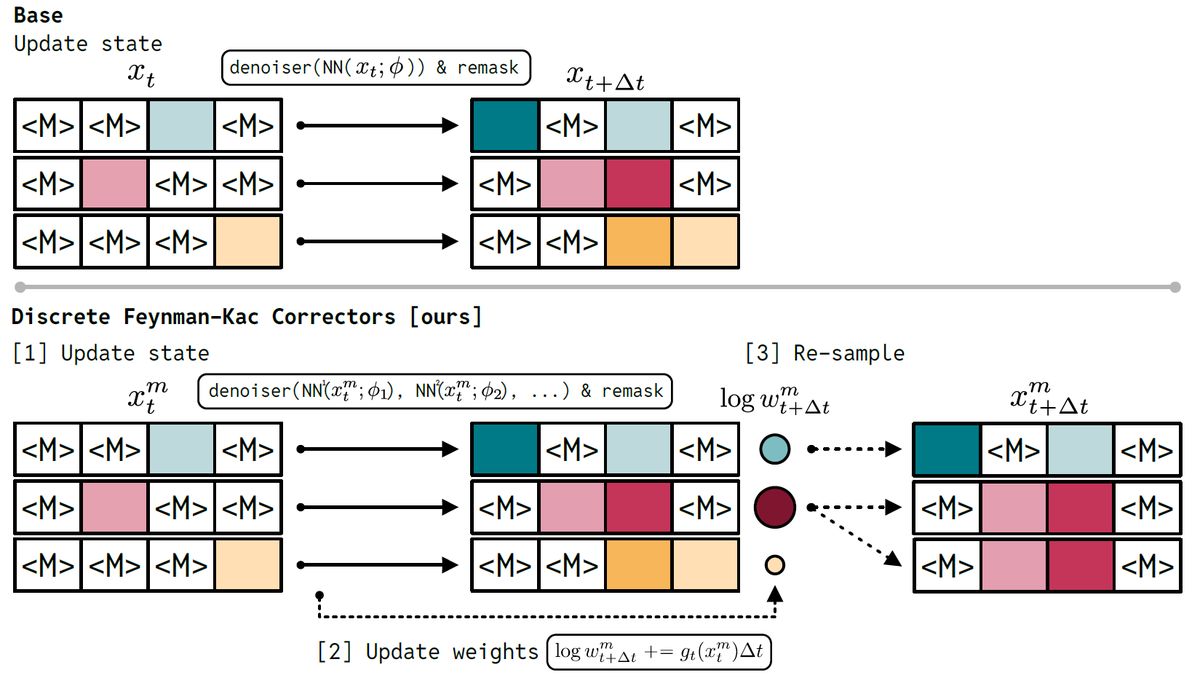
🚀New paper: Discrete Feynman-Kac Correctors!
It’s an inference time method that modifies discrete diffusion models via annealing, products, & reward-tilting. W/o training, it generalizes past the Ising critical temp. and boosts coding performance!
arxiv.org/abs/2601.10403 🧵👇
English
Artem Kushner retweetledi
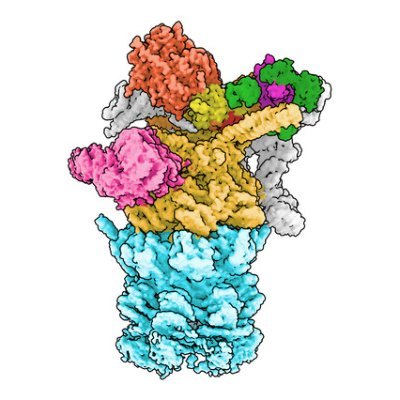
Have a look at our new structure of co translational folding in yeast. This is collaborative work initialized by the Rospert lab from the @UniFreiburg . Structural work has been done by the amazing Lorenz Grundmann.
IMP@IMPvienna
🧪Scientists from our Haselbach lab captured how proteins begin to fold as they’re being made. Using cryo-EM, they visualised chaperones guiding nascent proteins on the ribosome: nature.com/articles/s4146…
English
Artem Kushner retweetledi

For encoding, current data formats are designed for static models, limiting biological insights and AI potential. We're creating standardized, machine-readable, and human-interpretable models of dynamic structures. See more details: diffuse.science/posts/encoding/
English
Artem Kushner retweetledi

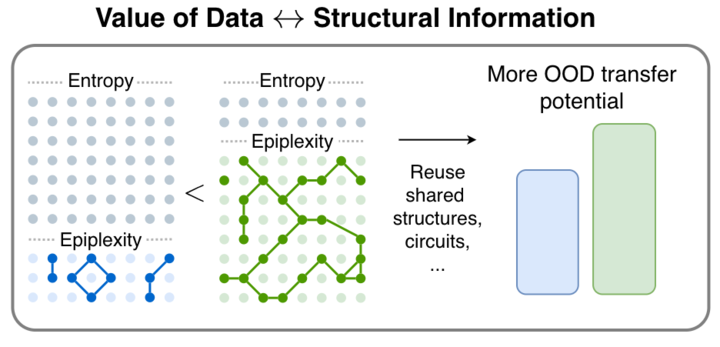
1/🧵 We are very excited to release our new paper! From Entropy to Epiplexity: Rethinking Information for Computationally Bounded Intelligence
arxiv.org/abs/2601.03220
with amazing team @ShikaiQiu @yidingjiang @Pavel_Izmailov @zicokolter @andrewgwils
English
Artem Kushner retweetledi

They provide a neat server visualizing all sorts of useful data about non-canonical amino acids and how they occur in PDB 👌 🤗
Paper: biorxiv.org/content/10.648…
Server: mondet.tuebingen.mpg.de
Very nice to easily see, e.g., how much training data of non-canonicals we have for each type and to quickly browse them!

English
Artem Kushner retweetledi

Can we apply gradient descent to discrete changes? In our new #SIGGRAPHAsia paper, we show that gradient descent can work on shape grammars, as in CAD and procedural modeling, but only if the grammars are designed correctly!
English
Artem Kushner retweetledi

AlphaFold predictions are valuable hypotheses and accelerate but do not replace experimental structure determination | Nature Methods nature.com/articles/s4159…
English
Artem Kushner retweetledi

Fast and interpretable quantification of biological shape heterogeneity via stratified Wasserstein kernel biorxiv.org/content/10.110… #biorxiv_bioinfo
English
Artem Kushner retweetledi

Don't worry if you missed MLSB! We recorded all of the talks!! 🕺💃
YouTube: @MLSBWorkshop/videos" target="_blank" rel="nofollow noopener">youtube.com/@MLSBWorkshop/…
Gina El Nesr@ginaelnesr
Absolutely blown away by the turn out at @workshopmlsb today. Thank you to the >600 people who came through the venue, and >150 that joined over Zoom throughout the day. Truly an exciting time to be innovating in ML for Structural Biology ♥️
English

