Su Datt Lam retweetledi
Su Datt Lam
87 posts


Su Datt Lam retweetledi
Su Datt Lam retweetledi

Have you ever wanted to design protein binders with ease? Today we present 𝑩𝒊𝒏𝒅𝑪𝒓𝒂𝒇𝒕, a user-friendly and open-source pipeline that allows to anyone to create protein binders de novo with high experimental success rates. @befcorreia @sokrypton
biorxiv.org/content/10.110…
English
Su Datt Lam retweetledi

Modeling protein-small molecule conformational ensembles with ChemNet @UWproteindesign
🚀 New preprint from David Baker!🚀
- ChemNet is a breakthrough deep learning model that accurately predicts the conformational ensembles of small molecules in complex with proteins.
- It outperforms existing methods like AlphaFold and RoseTTAFold by focusing on atomic-level accuracy, generating ensembles rather than single static structures.
- The key innovation is ChemNet’s stochastic nature, which allows it to model conformational heterogeneity, making it ideal for protein-small molecule docking.
- ChemNet generates diverse structures based on random initializations, providing deep insights into the flexibility of molecules at binding sites.
- The model is designed to assess the pre-organization of catalytic sites in enzymes, which is crucial for improving enzyme activity.
- In enzyme design applications, ChemNet-guided designs have achieved impressive success rates, including a retroaldolase with a kcat/KM of 11,000 M-1min-1, the highest pre-deep learning design rate for this reaction.
- The ability to rapidly generate conformational ensembles makes ChemNet a valuable tool for drug discovery and computational enzyme design.
- This system provides a faster, more generalizable alternative to computationally expensive methods like molecular dynamics simulations.
- ChemNet is expected to have broad applications in predicting protein-small molecule interactions, and guiding de novo design of enzymes and drug binders.
@ikalvet @gyurielee @alchemist_an @JustasDauparas
📜Paper: biorxiv.org/content/10.110…

English
Su Datt Lam retweetledi

Automated molecular optimization of a BRD4 inhibitor using #AlphaFold predicted structure with our new #GenerativeAI drug design method, DrugHIVE.
paper: pubs.acs.org/doi/10.1021/ac…
GitHub: github.com/jssweller/Drug…
@RemoRohs lab @qcb_usc. @JCIM_JCTC. #compchem #machinelearning
English
Su Datt Lam retweetledi

#AlphaFold3 is out, and good gravy is it a read. Congrats to the @GoogleDeepMind @IsomorphicLabs folks, some seriously compelling results especially for antibody-antigen complex predictions. Has got me thinking a lot!
Max Jaderberg@maxjaderberg
Super excited to be releasing AlphaFold 3 today, developed by @IsomorphicLabs and @GoogleDeepMind: our next generation AI model for predicting the biomolecular structures and interactions of proteins, DNA, RNA, small molecules, and more: bit.ly/44yfaCw 1/
English
Su Datt Lam retweetledi

AF3 server is LIVE! Just tried predicting complex with almost 5K amino acids.
TIP: you need to click "continue with google" to access the server (otherwise the "server" is grayed out). alphafoldserver.com


English
Su Datt Lam retweetledi

Thrilled to announce AlphaFold 3 which can predict the structures and interactions of nearly all of life’s molecules with state-of-the-art accuracy including proteins, DNA and RNA. Biology is a complex dynamical system so modeling interactions is crucial blog.google/technology/ai/…
English
Su Datt Lam retweetledi
Su Datt Lam retweetledi

Are there any privileged routes of acquisition of #driver alterations in #cancer?
Can they be used to #predict the outcome of treatments at diagnosis?
In our latest work, out in @NatureComms, we show that such routes do exist for most cancer types!
nature.com/articles/s4146…
1/6
English
Su Datt Lam retweetledi

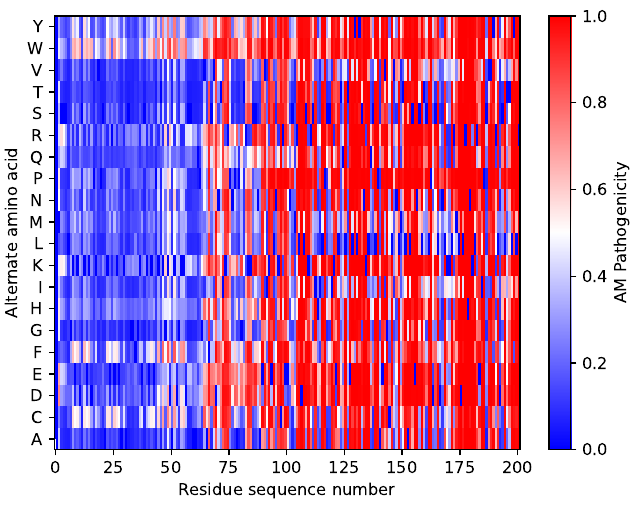
Want to know what a single-residue substitution does to your favorite protein?
Together with @TobiasRaisch we wrote a tool to easily visualize pathogenicity scores from the @GoogleDeepMind #AlphaMissense paper: github.com/MPI-Dortmund/p…
We hope you find it interesting :-)

English
Su Datt Lam retweetledi
Su Datt Lam retweetledi

The human genome is gradually unravelling its secrets 🎁
AlphaMissense model @ScienceMagazine: one more path lit up by deep learning in exploring the code of life 🧬
We now know with high confidence if 89% of ALL missense variants are benign or pathogenic
Key contributions🧵🧵

English
Su Datt Lam retweetledi

PICKLUSTER: A protein-interface clustering and analysis plug-in for UCSF ChimeraX biorxiv.org/cgi/content/sh… #biorxiv_bioinfo
English
Su Datt Lam retweetledi

What is an exon???
New paper w/ @ewjwallace and @RNA_julie in @CellGenomics : cell.com/cell-genomics/…
Cue 🧵1/8
English
Su Datt Lam retweetledi

And it's a wrap!! Thank you for coming! 😍🌷
Many thanks to @sudattlam @angmiayang @sndigregorio zetty and Prof Sheila Nathan, @griffiemma @FinlayM @pha4ge

Kuala Lumpur, Wilayah Persekutuan Kuala Lumpur 🇲🇾 English
Su Datt Lam retweetledi

And, hAMRonization 2.0 workshop is happening now 😃
@sudattlam @angmiayang @sndigregorio @griffiemma @FinlayM @pha4ge
Zetty giving opening remarks:

English
Su Datt Lam retweetledi

If you are looking for antibiotic summary cards:
International Society of Antimicrobial Chemotherap@ISAC_world
📢NEW Free Antibiotic Summary Cards Coinciding with #WorldHealthDay, the Alliance for the Prudent Use of Antibiotics (APUA) & ISAC have launched "Antibiotic cards" for some of the main antibiotics in use Download the cards here⬇ apua.org/antibiotic-car… #antimicrobialresistance
English
Su Datt Lam retweetledi

MIT Intro to Deep Learning - 2023 Lectures are Live
MIT Intro to Deep Learning is one of few concise deep learning courses on the web. The course quickly takes you to the foundations of deep learning, neural net architectures, and applications of DL.
introtodeeplearning.com


English
Su Datt Lam retweetledi

In a single long MD simulation we saw a loop change from a closed (red) to open (green) state, with many short "false openings" (yellow). In this pre-print, we characterize the loop residues that act as a conformational switch, and see that the motif occurs in other proteins too
D. E. Shaw Research@DEShawResearch
Our article “A conserved local structural motif controls the kinetics of PTP1B catalysis” has been released on @biorxivpreprint. doi.org/10.1101/2023.0…
English




