Chris Myers 💻🧬🧪 ☕ ℏ⁻³/N!∬exp(-βĤ) d𝓅 d𝓺
6.4K posts


Chris Myers 💻🧬🧪 ☕ ℏ⁻³/N!∬exp(-βĤ) d𝓅 d𝓺
@bayouphysicist
Professor Physics/Chemical Biophysics 💻🧬☕ 🌇 @fordhamnyc 🎓: @bcmhouston ⚕ @riceuniversity🦉 Formerly: @NYUniversity 🗽 @UmBiophys〽@UTMBhealth



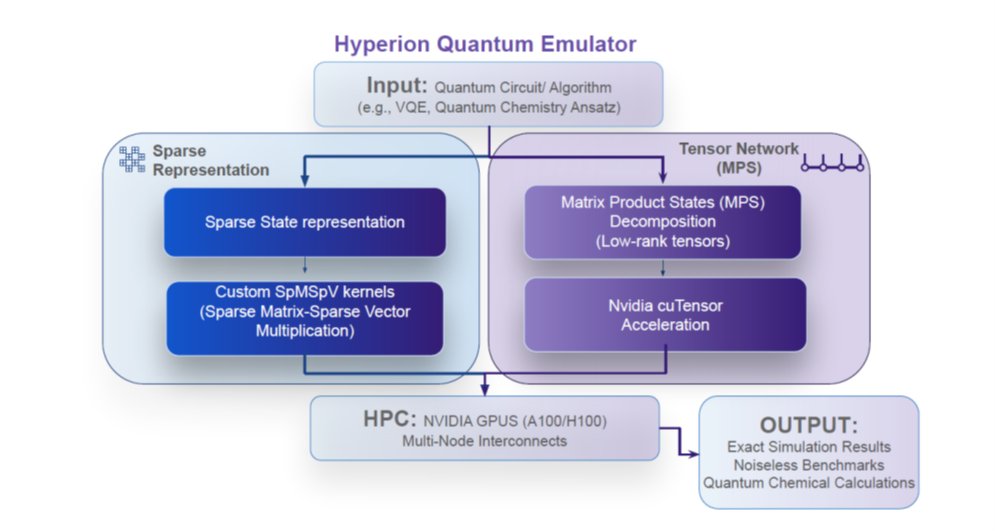
#compchem #machinelearning #quantumcomputing New preprint: "The Convergence Frontier: Integrating Machine Learning and High Performance Quantum Computing for Next-Generation Drug Discovery". @qubit_pharma arxiv.org/abs/2603.17790





We recently released a technical report highlighting how @IsomorphicLabs’ drug design engine is achieving a step change in how we model complex biological mechanisms, consistently outperforming industry benchmarks. Read our blog to discover how we’re bridging the gap between structure prediction and rational drug design. You can find the link in the comments.




@jk_rowling It will forever be amazing to me that people with medical degrees got in front of a camera and said that the effects of puberty blockers on children and adolescents are fully reversible. The horror.


#compchem #machinelearning First paper of the year, just out in J. Phys. Chem. Lett. ( @JPhysChem ): "Accelerating Molecular Dynamics Simulations with Foundation Neural Network Models using Multiple Time-Step and Distillation". Check the paper: pubs.acs.org/doi/full/10.10… (see also the updated preprint: arxiv.org/abs/2510.06562) We present a distilled multi-time-step (DMTS) strategy to accelerate molecular dynamics simulations using foundation neural network models. DMTS uses a RESPA-like formalism and a dual-level neural network where the target accurate potential is coupled to a simpler but faster model obtained via a distillation process. The approach conserves accuracy, preserving both static and dynamical properties. For those who read the initial preprint, the published version includes additional results leveraging active learning for enhanced stability. Such a strategy is applicable to any neural network potential and reduces their performance gap with classical force fields. Great work (and 1st paper!) by @comecattin. Kudos to the team: @Thomas__Ple, O. Adjoua, L. Lagardère and @NicolaiGouraud (@qubit_pharma). Funded by the @ERC_Research (project EMC2), supercomputing time @Genci_fr








We exposed the global far-right’s playbook at Munich, and how to stop them. Right on cue, Rubio beelined from his revisionist MAGA speech at MSC to boost Orbán in Hungary’s elections. We must wake up before it’s too late. A working class politics can defeat authoritarianism.






🇨🇳 China now generates 40% more electricity than the US and EU combined.