Dr. Neil Ashley retweetledi

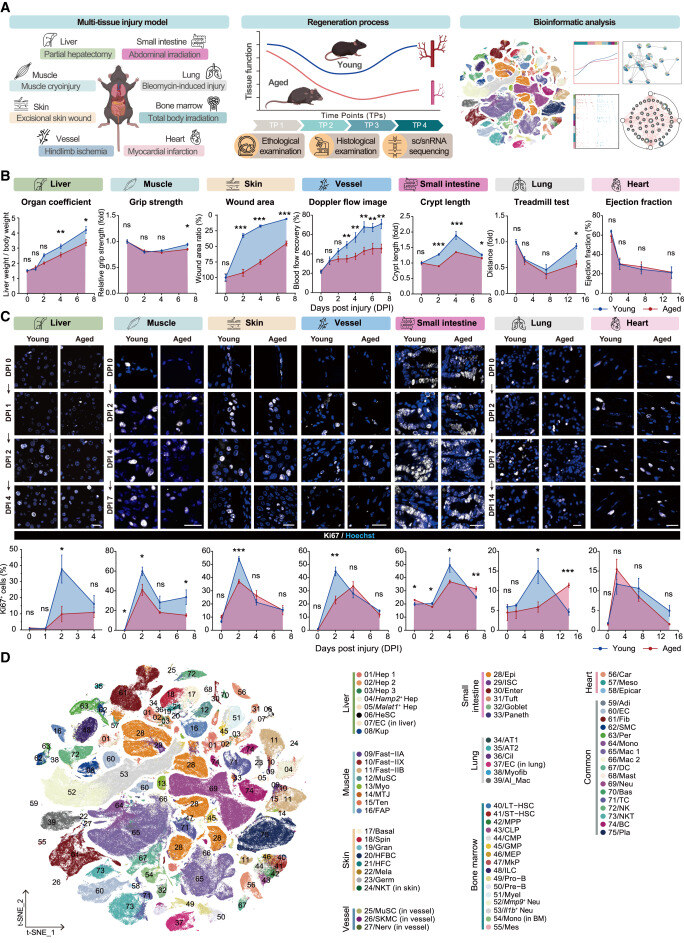
Decoding aging-dependent regenerative decline across tissues at single-cell resolution
hubs.li/Q037Z3bS0
Open stem cell research from Cell Stem Cell.
English
Dr. Neil Ashley
3.9K posts


@NotWIMM
Oxford based biotech scientist who likes single cell/spatial genomics #SINGLECELL #SPATIALGENOMICS #epigenetics