Gherman Novakovsky (слава Україні! 🇺🇦)
762 posts

Gherman Novakovsky (слава Україні! 🇺🇦)
@NovakovskyG
PhD, Illumina AI lab; interested in Deep Learning and genome regulation; also drawing, martial arts, guitar, and death metal! (he/him)

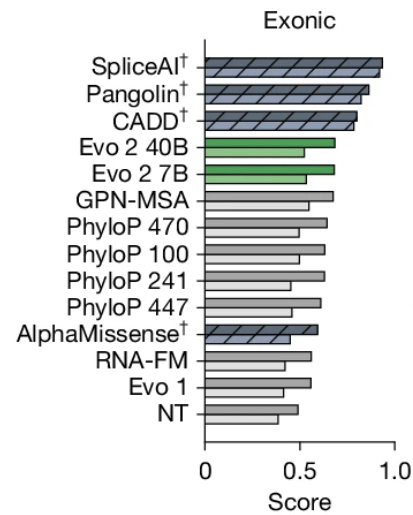

















*Easter egg alert* NOT in the published paper. We also benchmarked Evo 2 and while it did better than other gLMs (consistent that scale can improve gLMs), it still falls short of a basic CNN trained using one-hot sequences and far short of supervised SOTA x.com/pkoo562/status…

Genome annotation is falling behind how fast genomes can be assembled—but Johns Hopkins researchers @KuanHaoChao, @StevenSalzberg1, @elapertea, @alaina_shumate, @celinehohzm, & @alan_mayonnaise (+ @JakobHeinz9) have created a tool that can change that: cs.jhu.edu/news/solving-t…

Excited to share my first contribution here at Illumina! We developed PromoterAI, a deep neural network that accurately identifies non-coding promoter variants that disrupt gene expression.🧵 (1/)

We're thrilled to introduce PromoterAI — a tool for accurately identifying promoter variants that impact gene expression. 🧵 (1/)





