
Yajie Zhao, 赵亚杰
748 posts


@ZhaoDylan
Group Leader of Changping Laboratory & Visiting Scientist at @MRC_Epid & Human Geneticist🧬/ Photographer📷/Cat Enthusiast🐱



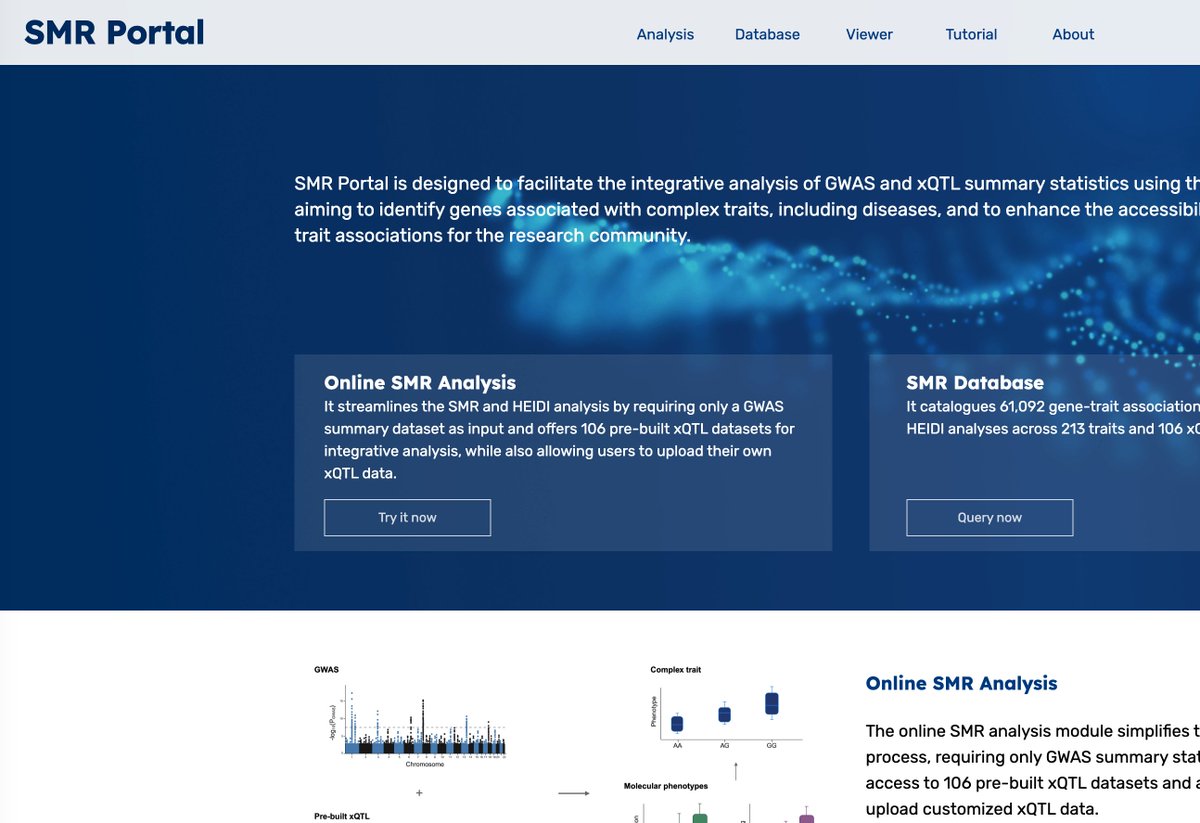
1/n New preprint! varTFBridge combines statistical association and fine-mapping with functional predictive models (VAR2TFBS, ABC-FP-Max, AlphaGenome) to trace common AND rare noncoding variants to the TFs and genes they affect. (1/) 📄 biorxiv.org/content/10.648…






It’s time to rethink Parkinson’s disease. Our work reframes PD as a disorder of the somato-cognitive action network (SCAN), and shows that normalization of SCAN connectivity represents a shared mechanism across diverse effective therapies. nature.com/articles/s4158… @ndosenbach @DavidRen555 @iamzhangvv @bttyeo @foxmdphd

