Jacob Corn
2.7K posts


Jacob Corn
@jcornlab
Genome editing, functional genomics, and cells figuring out how to eat themselves without dying. Professor of Genome Biology at ETH Zürich.
Zurich, Switzerland Katılım Şubat 2014
477 Takip Edilen5.7K Takipçiler

Cells need to repair genome editing induced damage, no matter how it’s made! This was a wonderful collaboration with @AllogeneTx.
Zacharias Kontarakis@xecoli
Our paper is out in Molecular Therapy – Nucleic Acids. We present #DisTALseq, a genome-wide method to profile TALEN #offtargets by tracking DNA repair factor recruitment at DSBs. Thank you all for this great collaboration #AllogeneTx #CornLab #GEML doi.org/10.1016/j.omtn…
English

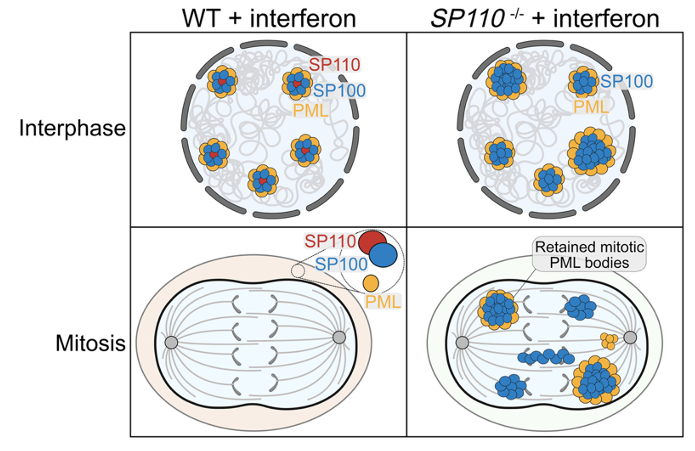
Proper organelle regulation during mitosis is a must, but how do membrane-less condensates like PML bodies behave during cell division? In work led by @eric_aird, we found the balance of speckled proteins SP110 & SP100 to be the key. doi.org/10.1038/s41556…
English

@ketanpatelimmo1 @karasuerman1 Excellent question. Maybe increased demand for NHEJ in HSCs plus more frequent overhanging ends vs blunt? No clear source of increased overhangs in HSCs so far as I know. My lab is more and more getting into tissue-specific etiologies of mutations in core repair factors.
English

@jcornlab @karasuerman1 But why should its loss lead to bone marrow failure?
English

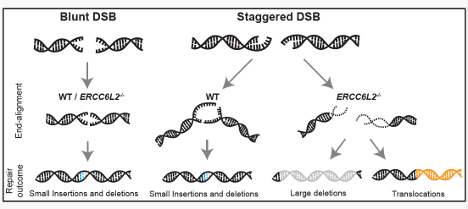
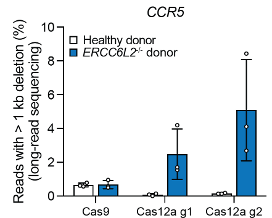
Many genome-editing tools work by cutting DNA - but the shape of the cut matters. Using #CRISPR screening, we show ERCC6L2 is specifically required to maintain #DNArepair fidelity at staggered DNA breaks, but not at blunt ends.
nature.com/articles/s4146…
English

@eric_aird @almuseben @SMSiegner @SPJacksonGroup @CejkaLab ERCC6L2 acts as a guardian during repair of staggered DSBs. This sheds light on ERCC6L2 mutations linked with BMF and leukemia and highlights caution for genome-editing approaches that generate DNA overhangs.
English

@eric_aird @almuseben @SMSiegner @SPJacksonGroup @CejkaLab What does ERCC6L2 actually do? It counteracts the activity of the MRN complex (MRE11–RAD50–NBS1), limiting excessive DNA end resection and promoting accurate end joining.
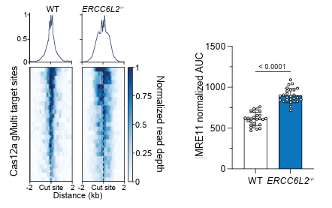
English

It’s been a fun first few week of my sabbatical. Many exciting meetings w/ @PeterFineran lab, learning new stuff and brainstorming ideas. And my own lab has been off to a crazy great start to the year. The early/late zoom meetings have me burning the candle at both ends…
English
Jacob Corn retweetledi

Happy to share that my postdoctoral work is now out in @ScienceMagazine. We show that the RNA-programmable bridge recombinase ISCro4 can insert, delete, or invert multi-kilobase DNA fragments at defined genomic sites in human cells.
Study link: science.org/doi/10.1126/sc…
CRISPR-Cas has transformed genome editing, but many diseases involve diverse patient-specific mutations within the same gene. A mutation-agnostic alternative is to insert a healthy gene copy at a defined genomic locus, but gene-sized, site-specific insertions remain a major challenge.
Main findings of our work:
(1) ISCro4 is highly active in human cells and can be delivered by plasmid or all-RNA formats
(2) Proof-of-concept programmable multi-kb insertions, deletions, and inversions
(3) Structural insights into the basis of enhanced ISCro4 activity
(4) Specificity and off-target characterisation
(5) A framework to support future development and adoption of bridge recombinases
The work was made possible through a close collaboration between the Jinek Lab, @schwanklab, and @jcornlab. I am deeply grateful to all of my co-authors for the team spirit, hard work, and dedication that went into this publication: @talasandris, Javier Fernández Carrera, @nicopmat, Lilly van de Venn, Charles Yeh, @p_kulcsar, @marquark, @YanikWeber, @SaskiaGerecke, Isabelle Harvey-Seutcheu, Dominic Mailänder, @MorPfl, and @ChrisChanez89. Special thanks to my supervisor, Prof. Martin Jinek, for his outstanding mentorship, and to Prof. Gerald Schwank and Prof. Jacob Corn for their generous support throughout. Finally, I would like to thank everyone in the Jinek lab for creating a supportive work environment, and @EMBO for funding my postdoctoral work.
In parallel, independent work led by @ntperry13, @SKonermann and @pdhsu also reported ISCro4 activity in human cells, reinforcing the robustness and momentum of this direction.
Please reach out if you would like to test the system or discuss potential applications. Relevant plasmids are now available via @Addgene.
English

This looks incredible! Cell systems can still be a bottleneck, but are cloning woes now a thing of the past?
Thomas Norman@thenormanlab
New preprint on technologies to scale up CRISPR screens. We use them to map 665,856 pairwise genetic perturbations and outline a path to comprehensive interaction mapping in human cells. We also introduce an approach for cloning lentiviral libraries with billions of elements.
English


AI reviewers are here — we are not ready nature.com/articles/d4158…
English
Jacob Corn retweetledi

1/ New preprint out! We show that PROTAC-induced ubiquitination can bypass canonical ERAD to degrade ER membrane proteins. Wonderful collaboration w/ @DanNomura and huge credit to grad student superstar Sydney Tomlinson!
biorxiv.org/content/10.110…
English