
Takashi Fukaya
873 posts

@FukayaLab
Institute for Quantitative Biosciences, UTokyo. We study the mechanism of transcriptional regulation in the Drosophila embryo.



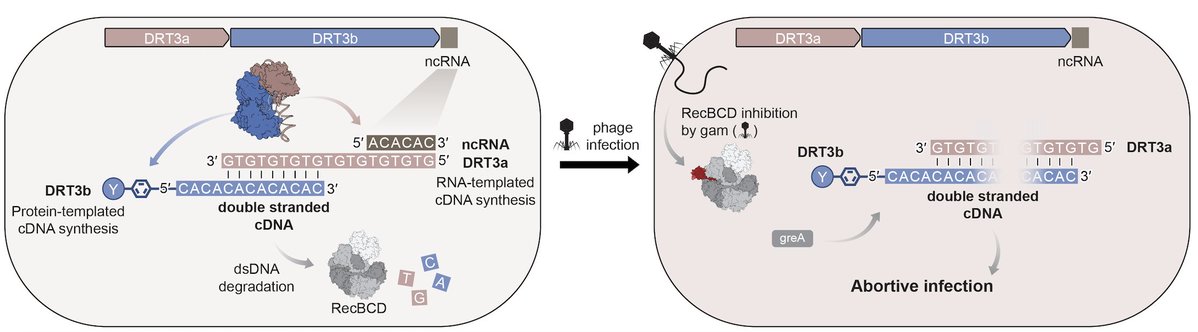
A preprint‼️that's bound to ruffle some 🪶 "Widespread DNA off-targeting confounds studies of RNA chromatin occupancy" led by our @MicahGoldrich and Louis Delhaye from @pieter_mestdagh. TL;DR we show that many of lncRNA chromatin occupancy maps are flawed🧵biorxiv.org/content/10.110…












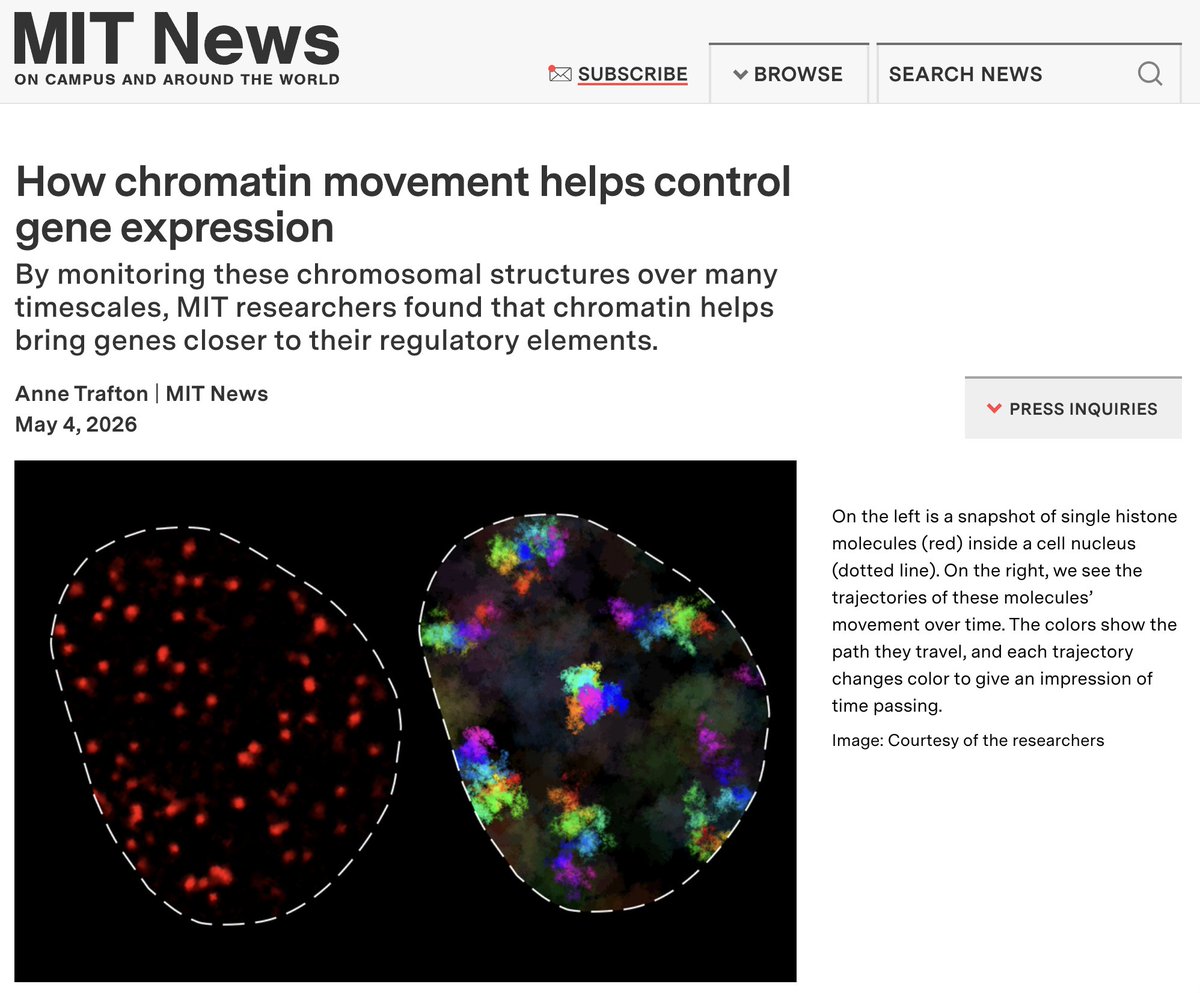
(1/13) Thread on @mazzocca_matteo @DomenicNarducci @SGrosseHolz @_jessematthias new preprint Q: how does chromatin move? Using MINFLUX, SPT & SRLCI, we track chromatin dynamics across 7 orders of magnitude in time to provide some answers biorxiv.org/content/10.110…


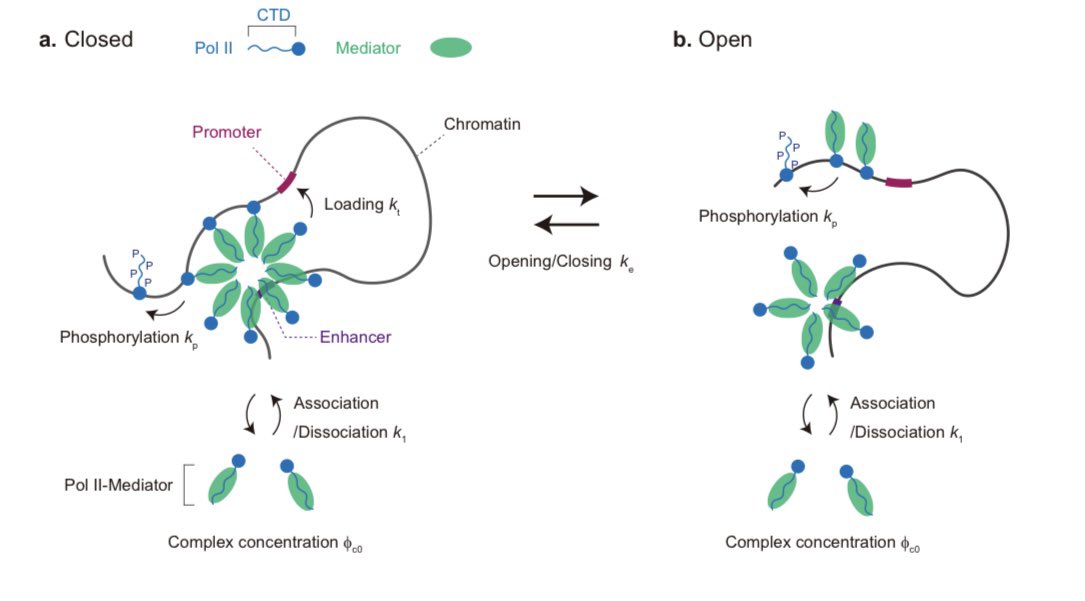

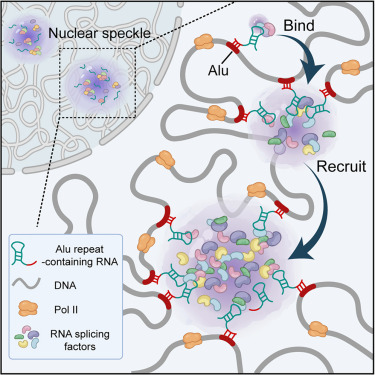
New preprint alert! Together with Tetsuya Yamamoto at Hokkaido University, we propose a “polymer micelle” model of transcriptional bursting. biorxiv.org/content/10.648…




