Nick Edwards
1.8K posts

Nick Edwards
@Nick___Edwards
Autonomous science. Founder and CEO at Potato (@readysetpotato). Former neuro at Brown, NIH, UCSD.


Humanity, created by God in all its grandeur, is today facing a pivotal choice: either to construct a new Tower of Babel or to build the city in which God and humanity dwell together. In Jesus Christ, this humanity in its grandeur becomes the Way, the Truth and the Life, opening the path for each of us to grow toward fullness. #MagnificaHumanitas vatican.va/content/leo-xi…

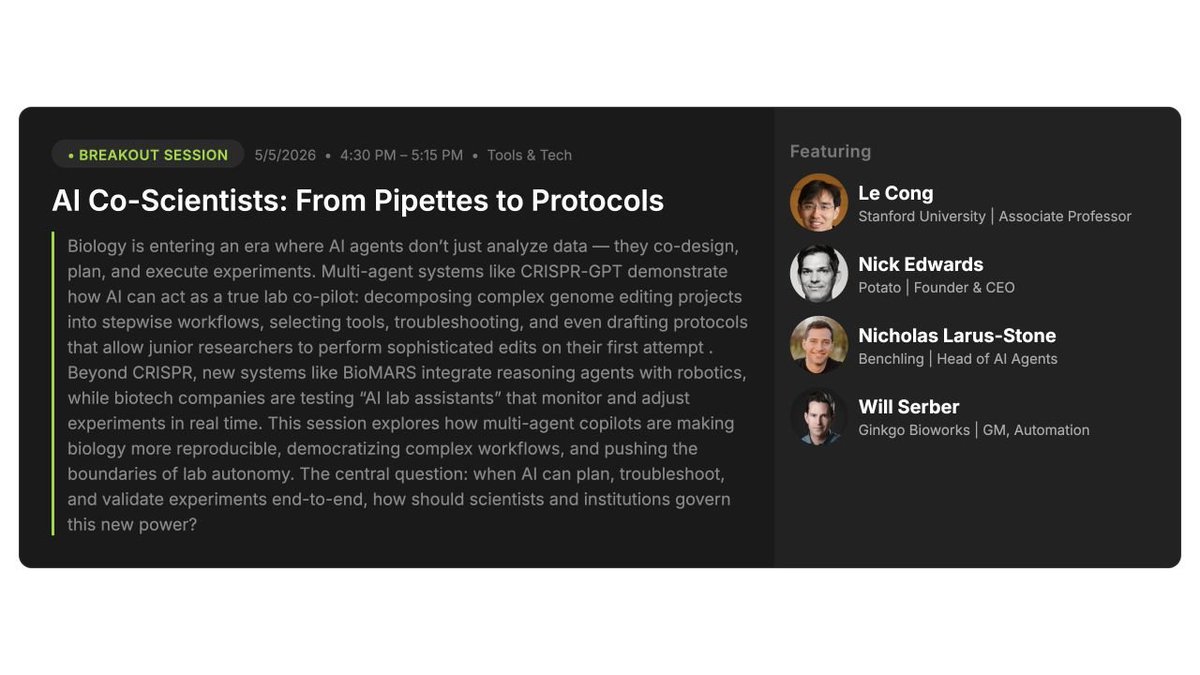










Alex Karnal (@alex_karnal) is the most talented bio and healthcare investor I've ever met. He's spent 20 years in the industry and says 2025 was the single most exciting year he's seen. The start of a once-in-a-lifetime, trillion-dollar revolution in public health. He explains how few people realize we already have the medicines to prevent our deadliest diseases. The problem is that almost no one takes them. There's a population of people born with a mutation that means their bodies don't produce a protein called PCSK9. Their lifetime risk of cardiovascular disease is 88% lower than yours. Pharma turned that genetic advantage into a drug. It's been approved for years, but the number of people taking it is still vanishingly small. Partly because high cholesterol is a silent killer. You feel nothing, right up until you have a heart attack. And partly because the health system makes it punishingly hard to stay on a preventive drug like a PCSK9 inhibitor. In other words, the medicine works, but the system around it doesn't. That's what's starting to change, and in this episode, Alex explains why. We discuss the "health stack" he believes can add a decade to most lives, why oral GLP-1s are breaking every adoption record in pharma, peptides and citizen pharmacology, and what AI is doing to drug discovery. I wish I had an "Alex" for every interesting topic. We've been having versions of this conversation for over five years, and every single one is as clear and as useful as this one. Enjoy! Timestamps: 0:00 Intro 1:00 The State of Modern Medicine 5:00 Designing the Modern Health Stack 12:17 The GLP-1 Inflection Point 19:18 The Biological Mechanisms of GLP-1 30:36 Overcoming Frictions in Healthcare 34:19 Cardiovascular Disease 44:04 Addressing Alzheimer's 47:04 The Future of Cancer 57:33 Drug Discovery 1:05:25 AI and Scientific Super Intelligence 1:14:40 Citizen Pharmacology and the Peptide Movement 1:18:13 Background and Career Journey 1:31:09 Braidwell's Investment Approach 1:33:30 The Kindest Thing