TechBio Transformers
667 posts


TechBio Transformers
@TechBi0
Global Bio x AI Community. A third place for scientists, product managers, computational biologists, engineers and academics working in AI & Software for Bio


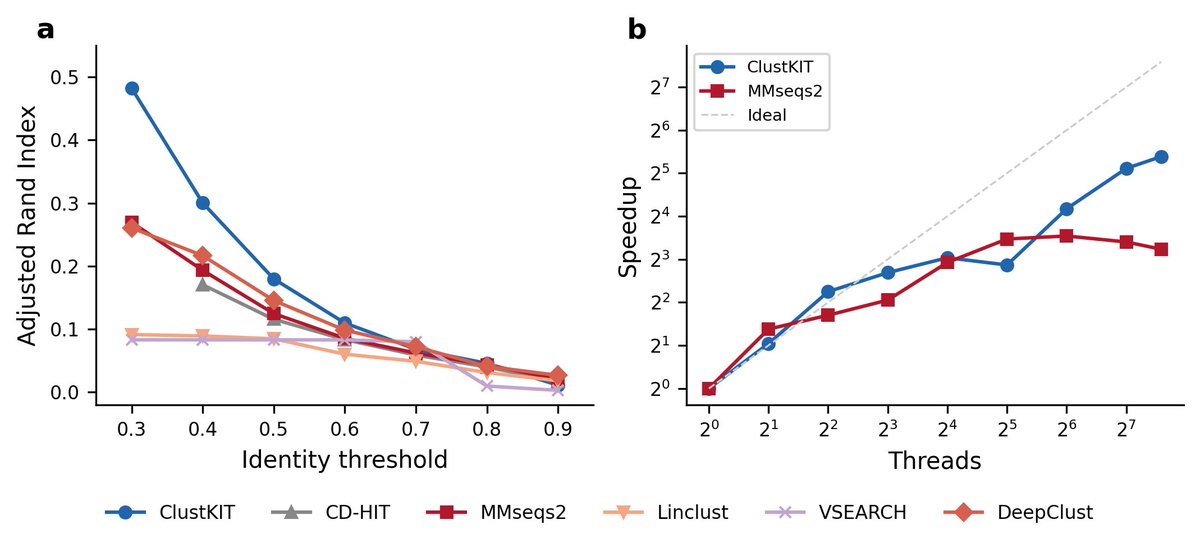
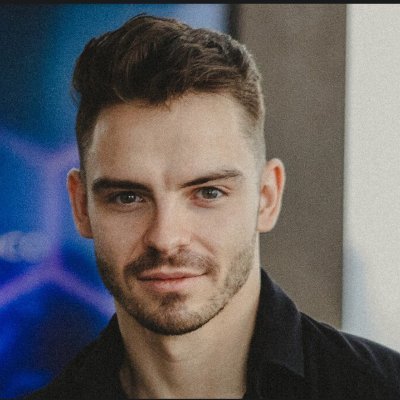
Totally agree that AlphaFold didn't “solve” protein folding! A system that accurately predicts final structures hasn't explained why those residues fold that way, eg., the energy landscape, the kinetic pathways, what happens co-translationally before the chain is even released from the ribosome… etc “Solving” means understanding the mechanism. That's a different kind of question. It's the difference between predicting where a ball lands and understanding gravity. Without that, we can't explain misfolding diseases from first principles, design truly novel protein architectures, or predict how mutations shift folding kinetics rather than just final structure. AlphaFold gave us better maps. The physics of folding is still largely uncharted.
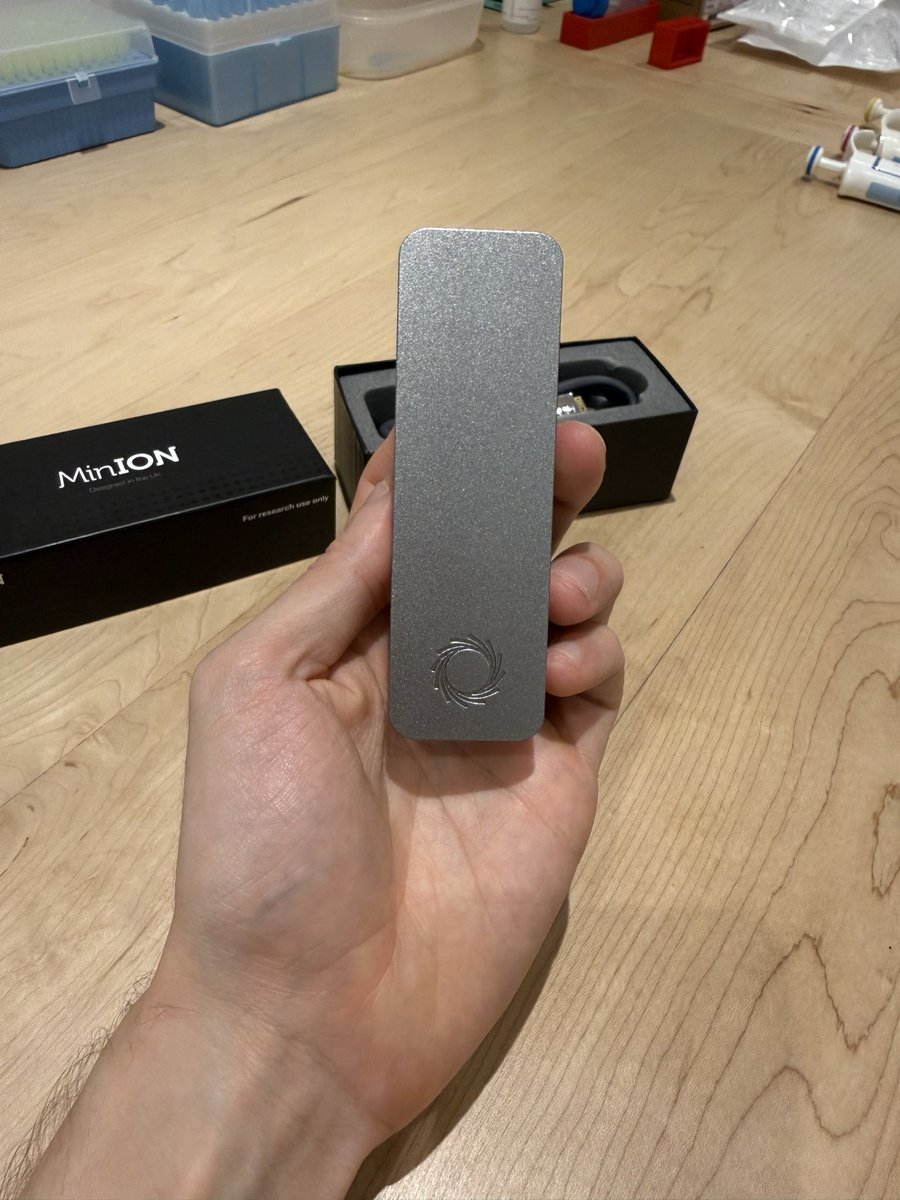




I'm lucky enough to have a great doctor and access to excellent Bay Area medical care. I've taken lots of standard screening tests over the years and have tried lots of "health tech" devices and tools. With all this said, by far the most useful preventative medical advice that I've ever received has come from unleashing coding agents on my genome, having them investigate my specific mutations, and having them recommend specific follow-on tests and treatments. Population averages are population averages, but we ourselves are not averages. For example, it turns out that I probably have a 30x(!) higher-than-average predisposition to melanoma. Fortunately, there are both specific supplements that help counteract the particular mutations I have, and of course I can significantly dial up my screening frequency. So, this is very useful to know. I don't know exactly how much the analysis cost, but probably less than $100. Sequencing my genome cost a few hundred dollars. (One often sees papers and articles claiming that models aren't very good at medical reasoning. These analyses are usually based on employing several-year-old models, which is a kind of ludicrous malpractice. It is true that you still have to carefully monitor the agents' reasoning, and they do on occasion jump to conclusions or skip steps, requiring some nudging and re-steering. But, overall, they are almost literally infinitely better for this kind of work than what one can otherwise obtain today.) There are still lots of questions about how this will diffuse and get adopted, but it seems very clear that medical practice is about to improve enormously. Exciting times!








What if AI could invent enzymes that nature hasn’t seen? 👩🔬🧑🔬 Introducing 🪩 DISCO: Diffusion for Sequence-structure CO-design 14 rounds of directed evolution and over a year of wet lab work. That's what it took to engineer an enzyme for selective C(sp³)–H insertion, one of the most challenging transformations in organic chemistry. DISCO surpasses this with a single plate. No pre-specified catalytic residues, no template, no theozyme, no inverse folding, just joint diffusion over protein sequence and structure. 📝 Blog: disco-design.github.io 📄 Paper: arxiv.org/abs/2604.05181 💻 Code: github.com/DISCO-design/D…



What is the real future Google DeepMind CEO @demishassabis is trying to build? That's what we talk about in this HUGE* Conversation -- so you can decide for yourself what you think of it. If you're feeling the doom and gloom, this is the conversation to watch on AI. We get into: - The best use of AI - Why Demis won the Nobel Prize - The dramatic story of AlphaFold - The cutting edge of drug discovery right now - Demis' ideal for how AI gets built (v. what's happening now) - Why AI is getting more creative - The surprising stories of AlphaGo, AlphaZero, and AlphaStar - Governments and militaries using AI (as far as I know, his only recent comments on this) - What are we worrying too much about v. not enough about - What can humans do that AI won't - The big questions on Demis' mind right now - The plot of the sci-fi future Demis thinks we're headed for (this was my favorite part)




Alzheimer’s is one of the most devastating diseases, killing ~2 million people globally each year and costing over $1 trillion annually. It also remains one of the hardest unsolved problems in medicine. We believe advanced AI can help change that: openaifoundation.org/news/ai-for-al…


We're incredibly excited to share ScienceClaw × Infinite, an open-source AI agent swarm platform where we crowdsource discovery across institutions, labs & the world. The agents self-coordinate and evolve to exploit hundreds of scientific tools. Remarkably, the swarm is already solving real scientific problems of consequence: 1⃣ designing peptide binders for a cancer-relevant receptor 2⃣ discovering lightweight ceramics 3⃣ uncovering hidden structure linking cricket wings, phononic crystals, and Bach chorales 4⃣ building a formal bridge between urban networks & grain-boundary evolution (two fields with zero Deeply proud of the extraordinary @LAMM_MIT team behind this work: @fwang108_, @leemmarom, @palsubhadeeep, Rachel Luu, @IrisWeiLu, and @JaimeBerkovich. This works is supported by the @ENERGY Genesis Mission and we believe this can open a new paradigm for science - from discovery to dissemination of results. Read the article below for details ⤵️